Search Count: 8
All
Selected
 |
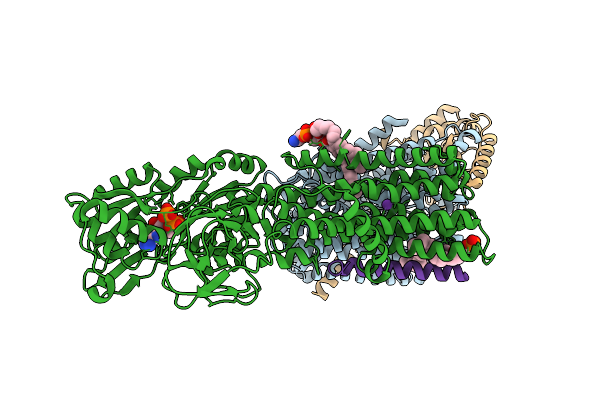
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1 State In Lipid Nanodisc
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.34 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1-Atp State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, ATP |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, MG |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.58 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2 State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
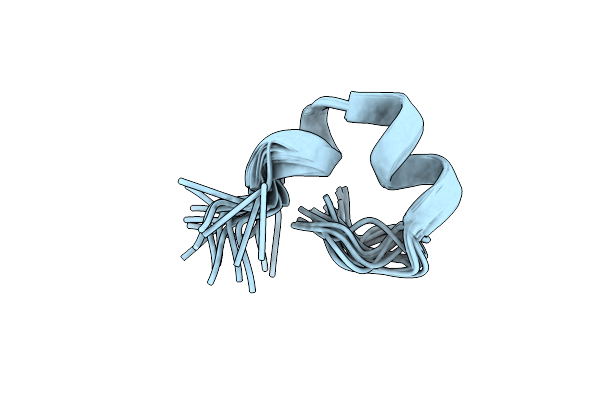
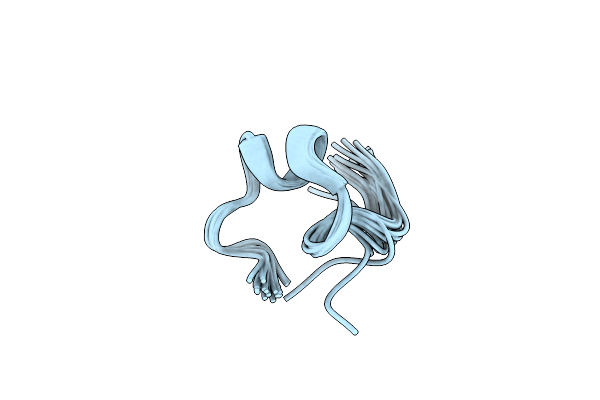
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2021-06-02 Classification: MEMBRANE PROTEIN |
 |
 |

