Search Count: 763
All
Selected
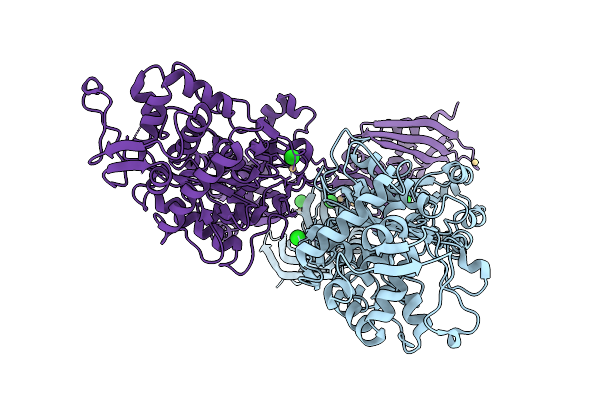 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
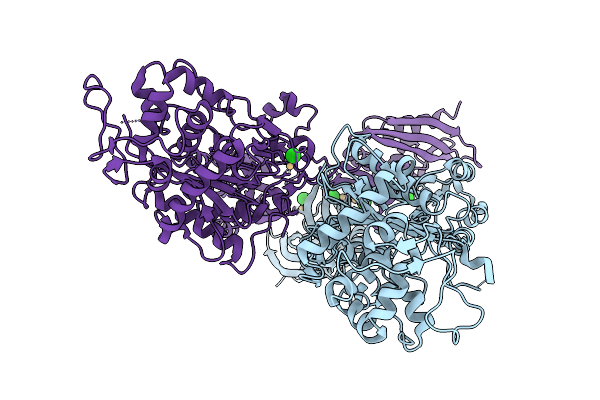 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: CD, CL |
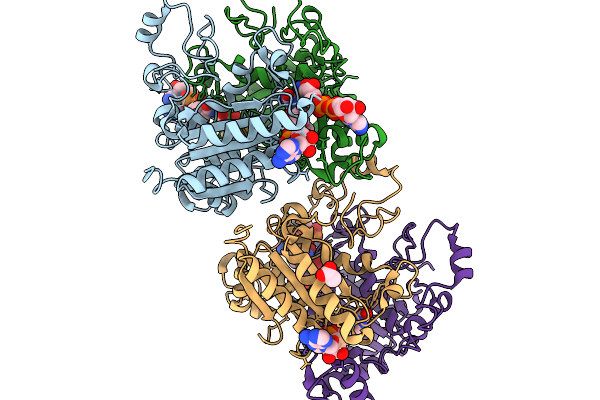 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: NAD, FAD, ACT |
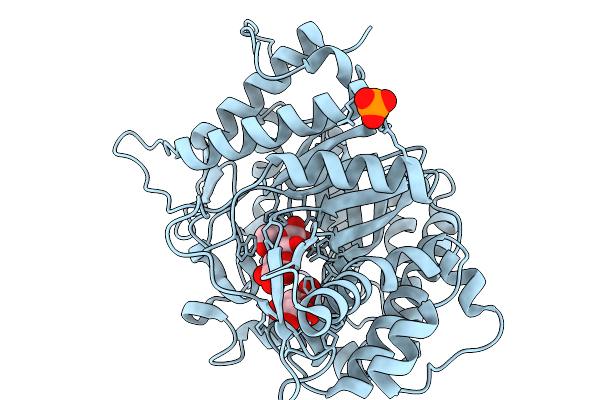 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, BGC, PO4 |
 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
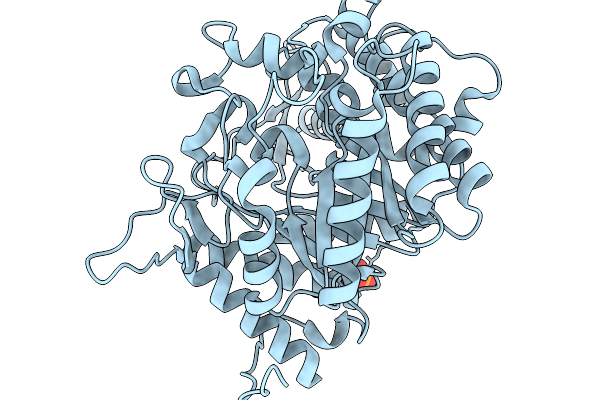 |
Crystal Structure Of Glucose Tolerant Beta-Glucosidase Unbgl1 Mutant (C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: CL, PO4 |
 |
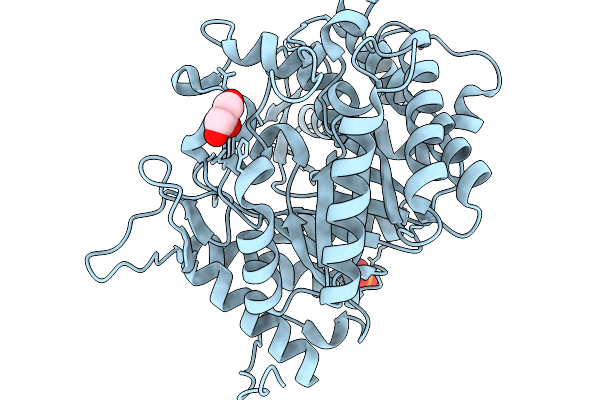
Crystal Structure Of Glucsoe Bound Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
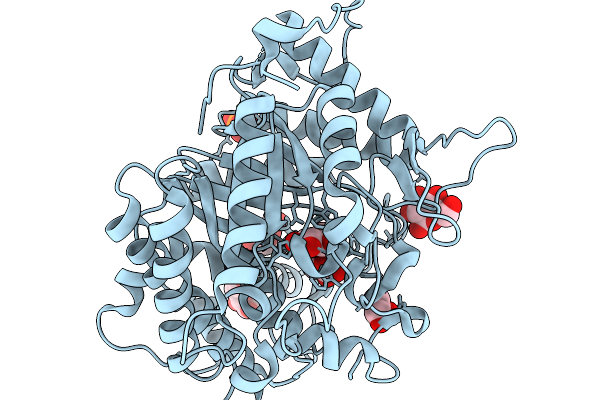
Crystal Structure Of Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, GOL, CL |
 |
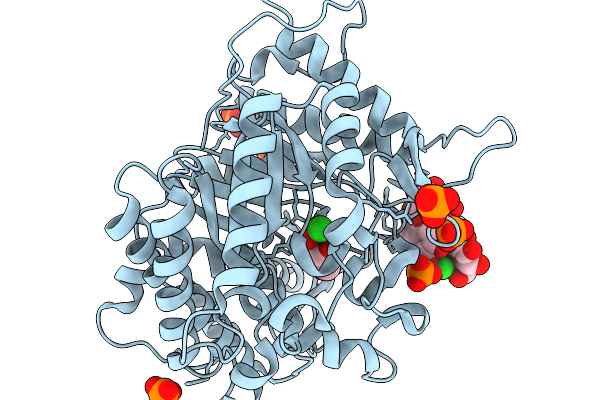
Crystal Structure Of Glucose Bound Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, GOL, PO4 |
 |
Crystal Structure Of Cellobiose And Glucose Bound Glucose Toleranant Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4, CL |
 |
High Resolution Crystal Strucutre Of Highly Glucose Tolerant Gh1 Beta-Glucosidase (Unbgl1_C188V_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, GOL |
 |
Crystal Structure Of Glucose Bound Highly Glucose Tolerant Gh1 Beta-Glcosidase Mutant (Unbgl1_C188V_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GS1, PO4, NA |
 |
Crystal Structure Of Glucose Bound Covalent Intermediate Of Gh1 Beta-Glucosidase (Unbgl1)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
High Resolution Crystal Structure Of Gh1 Beta-Glucosidase From Soil Metagenome (Unbgl1)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, CL, GOL |
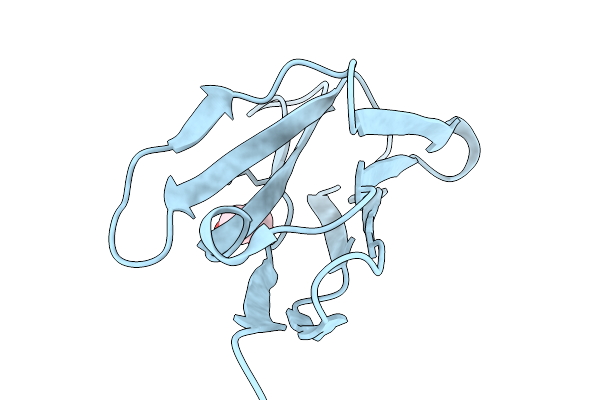 |
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM Ligands: EDO |
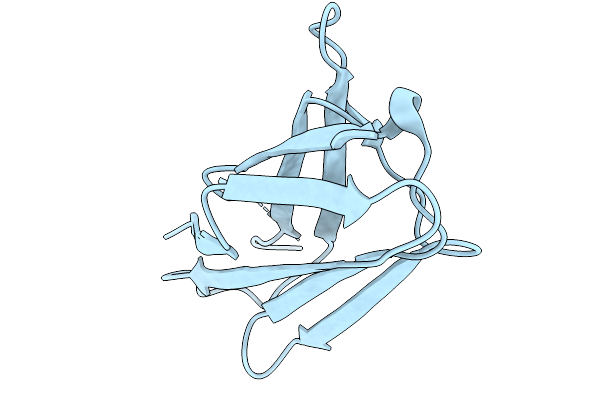 |
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM |
 |
Group Ii Intron Assembly Intermediate Domain 1, 2, 3 And 4 "Fully Open" State
Organism: Oceanobacillus iheyensis
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: RNA |
 |
Xfel Structure Of Hnqo1 Mixed With Nadh In An Extended Orientation At 0.3 S
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: NAD, FAD, ACT |
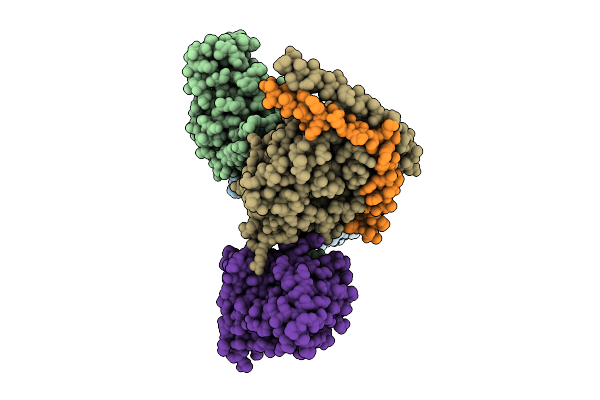 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: SIGNALING PROTEIN |

