Search Count: 2,775
All
Selected
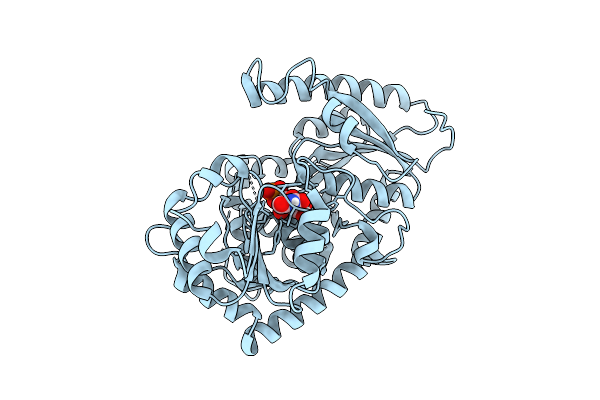 |
Crystal Structure Of The Glycosyltransferase Qsfuct From Quillaja Saponaria
Organism: Quillaja saponaria
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP |
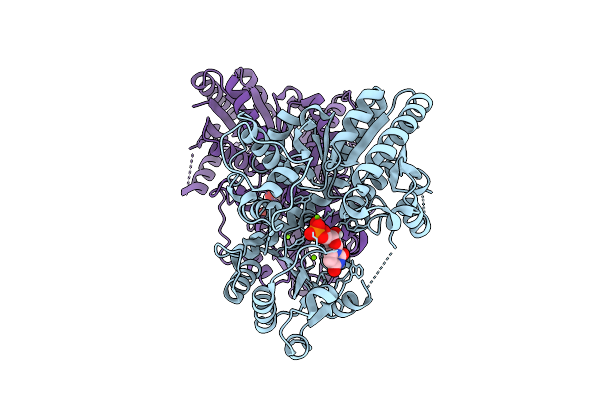 |
Crystal Structure Of The Glycosyltransferase Svfuct From Saponaria Vaccaria
Organism: Gypsophila vaccaria
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP, MG |
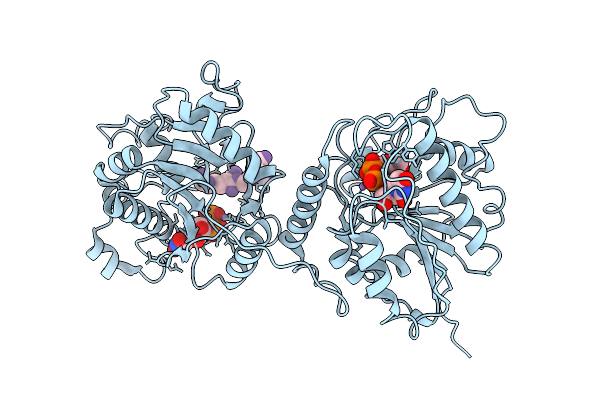 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN, UDP |
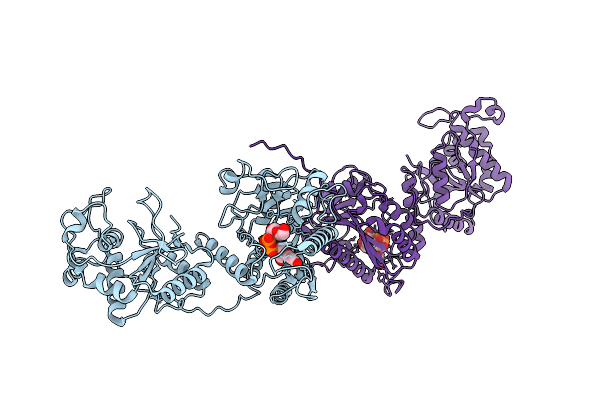 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
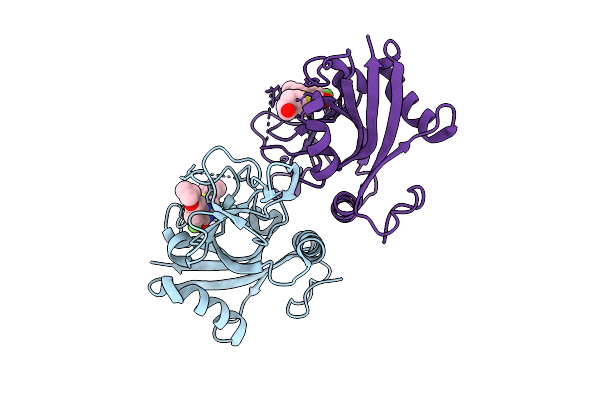 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: XBA |
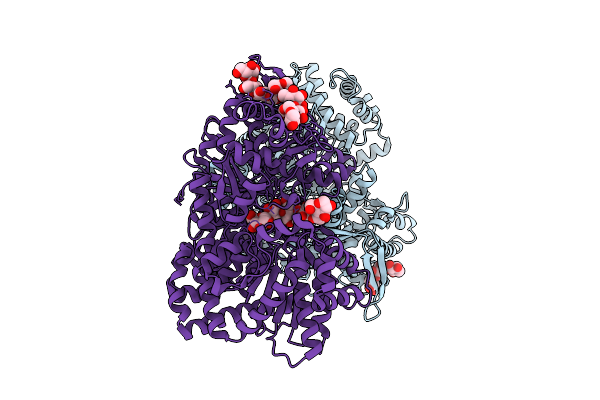 |
Carbohydrate-Bound Structure Of Alpha-Glucan Phosphorylase From Crocosphaera Subtropica Atcc 51142
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: TRANSFERASE |
 |
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: MG, TRT, A1BYH |
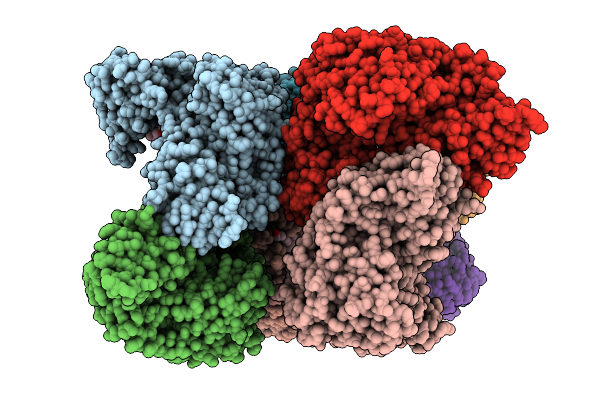 |
Organism: Nocardia brasiliensis
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: MG, TON, A1BYH, TRT |
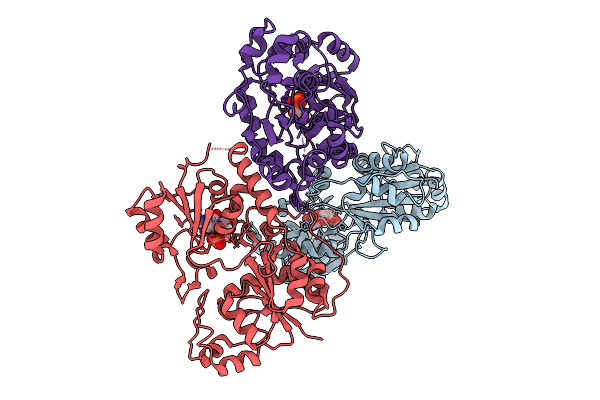 |
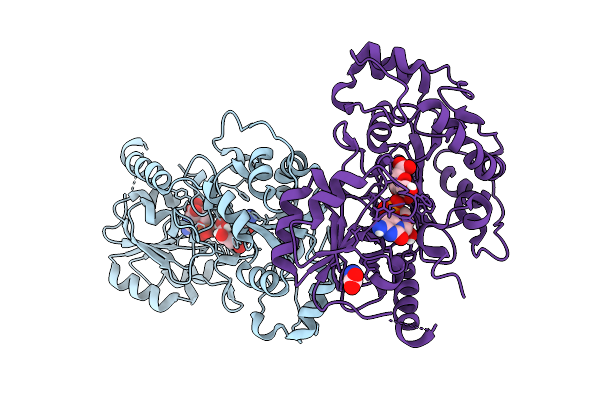
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: C5P, CL |
 |
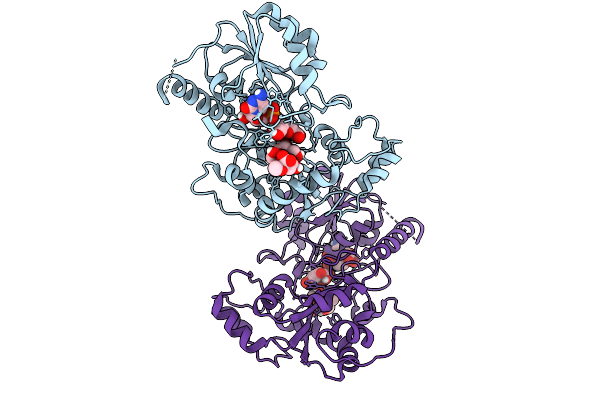
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: C5P, KDO, GLY, CL |
 |
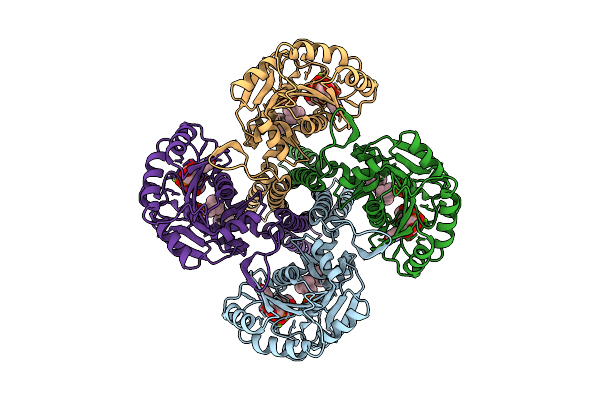
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: C5P, KDO, CL |
 |
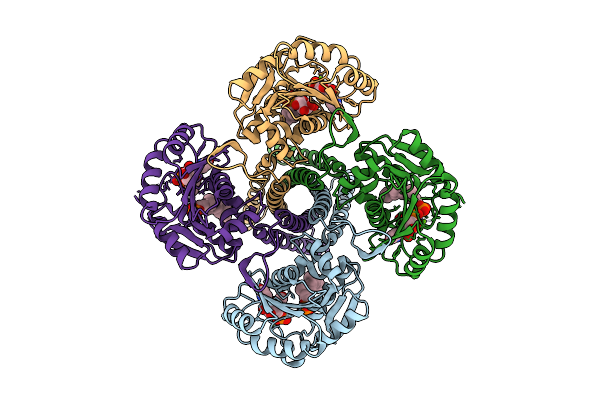
Cryo-Em Structure Of The Glycosyltransferase Gtrb In The Substrate-Bound State
Organism: Synechocystis sp. pcc 6803 substr. kazusa
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN Ligands: UPG, 5TR, MG |
 |
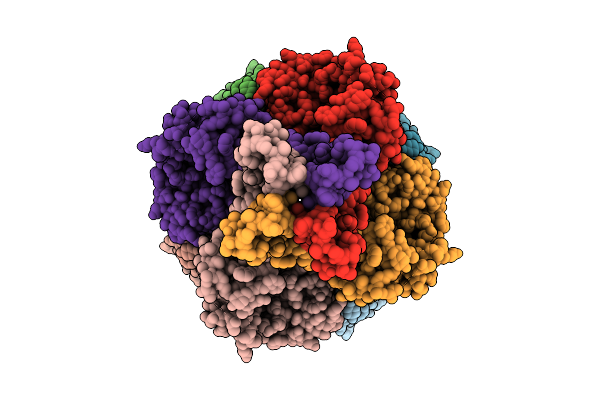
Cryo-Em Structure Of The Glycosyltransferase Gtrb In The Pre-Catalysis And Product-Bound State
Organism: Synechocystis sp. pcc 6803 substr. kazusa
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN Ligands: UPG, 5TR, MG, UDP, A1B94 |
 |
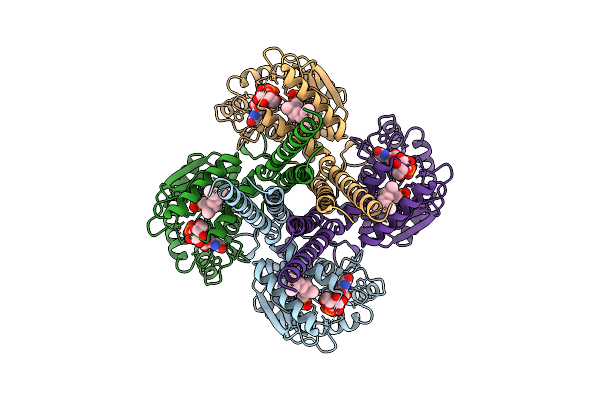
Cryo-Em Structure Of The Glycosyltransferase Gtrb In The Apo State (Octamer Volume)
Organism: Synechocystis sp. pcc 6803 substr. kazusa
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
 |
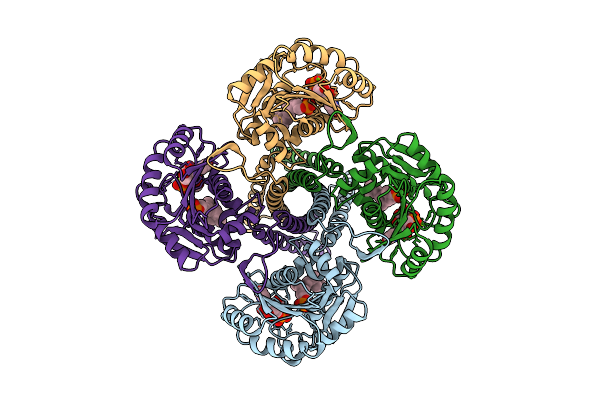
Organism: Synechocystis sp. pcc 6803 substr. kazusa
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN Ligands: UPG, 5TR, MG |
 |
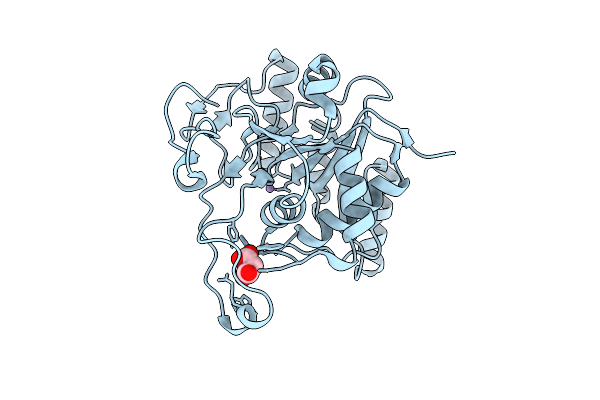
Cryo-Em Structure Of The Glycosyltransferase Gtrb In The Pre-Intermediate State
Organism: Synechocystis sp. pcc 6803 substr. kazusa
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN Ligands: UPG, 5TR, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-12-10 Classification: SUGAR BINDING PROTEIN Ligands: MN, MLI |
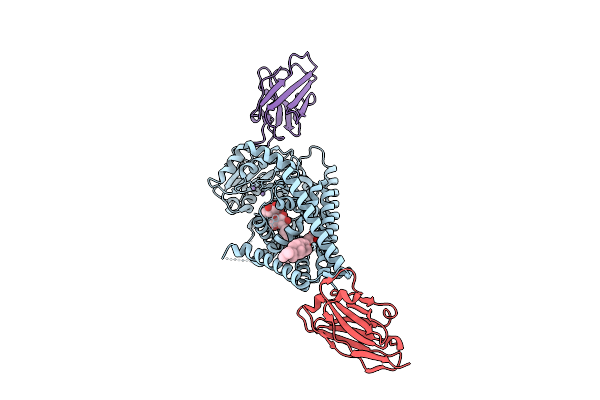 |
Organism: Paramecium bursaria chlorella virus cz-2, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: TRANSFERASE/IMMUNE SYSTEM Ligands: Y01, MN, LMT |
 |
Organism: Paramecium bursaria chlorella virus cz-2, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: TRANSFERASE/IMMUNE SYSTEM Ligands: UGA, MN, Y01 |

