Search Count: 13
All
Selected
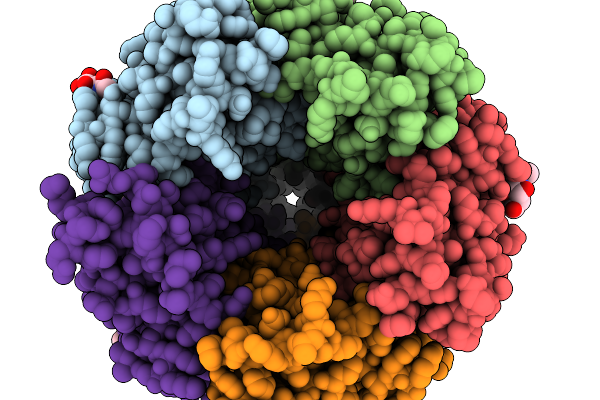 |
Alpha1/Beta Heteromeric Glycine Receptor In The Presence Of 0.200 Mm Strychnine And 500 Nm Ivermectin
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Resolution:2.86 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: SY9, NAG |
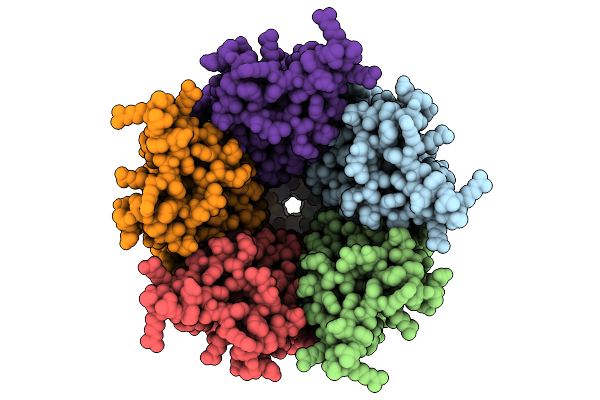 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiiii Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: PEE |
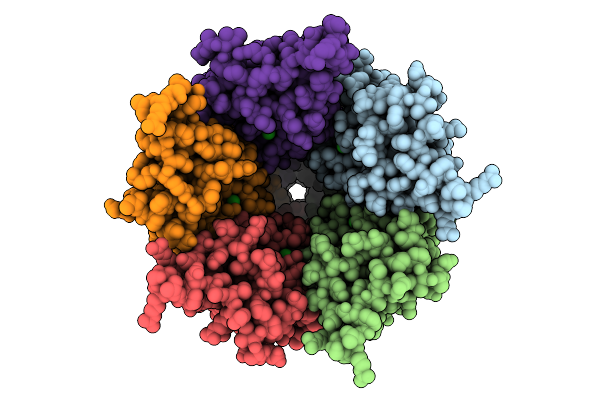 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiiio Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
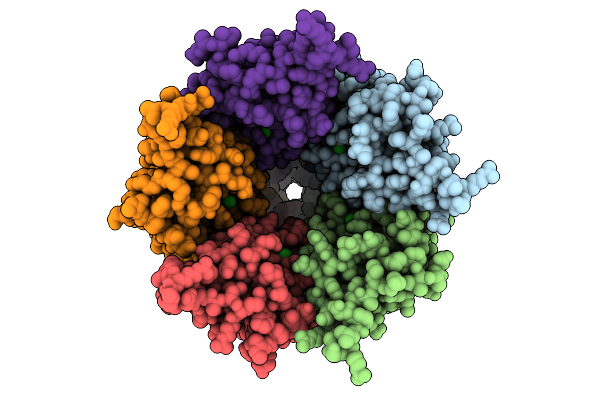 |
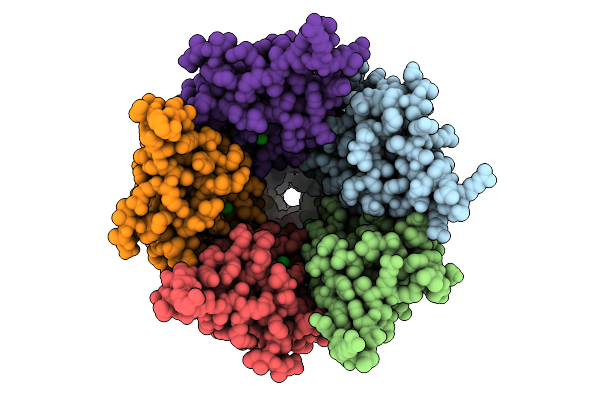
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiioo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
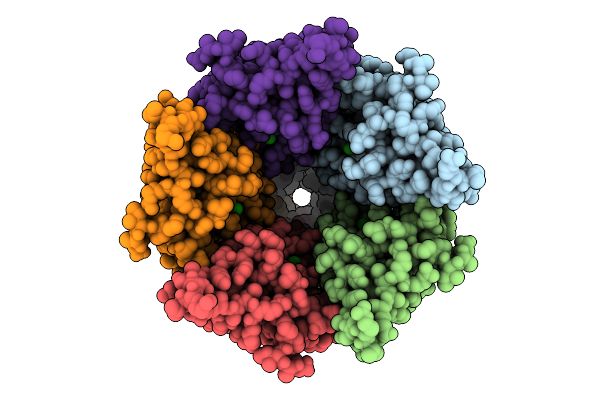
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
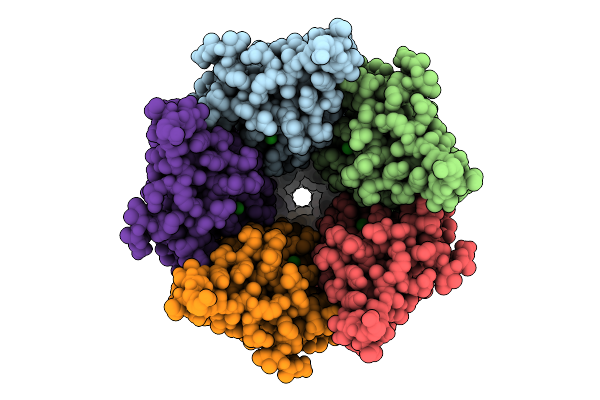
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Ioooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
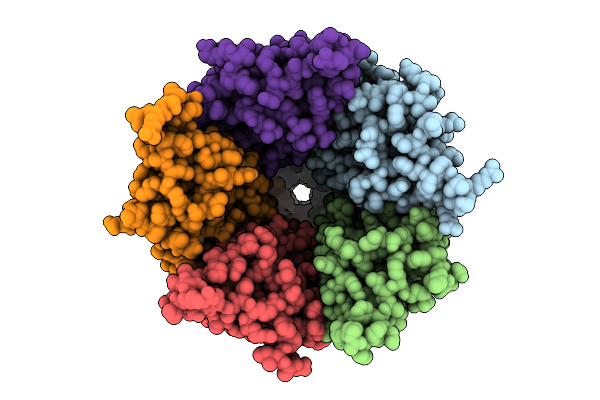
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Ooooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
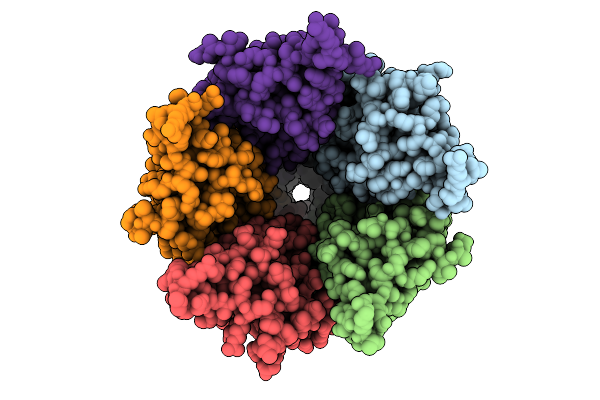
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iioio Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.45 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: PEE |
 |
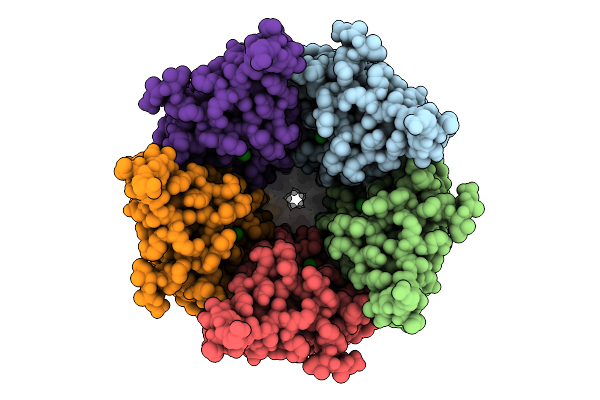
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Ioioo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: PEE |
 |
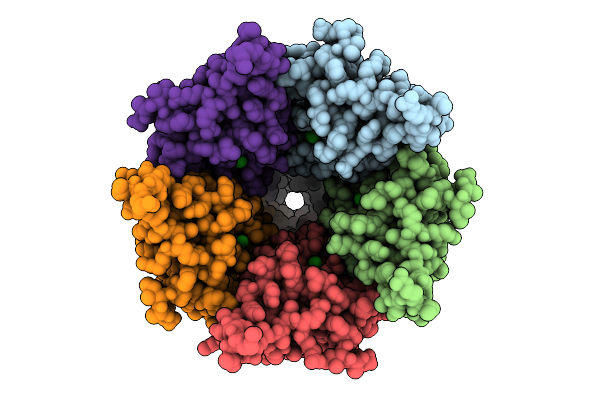
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic F116A Mutant At Ph 2.5
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:2.14 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
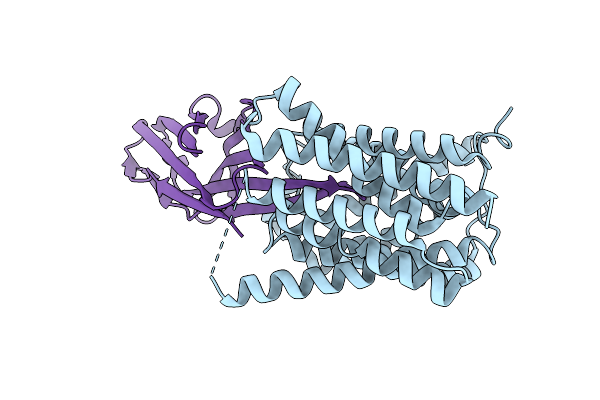
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic Y251A Mutant At Ph 2.5 In Ioooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
 |
Cryo-Em Structure Of Candida Albicans Vrg4 Bound To An Inhibitory Nanobody.
Organism: Lama glama, Candida albicans
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-09-03 Classification: TRANSPORT PROTEIN |
 |
Cryo-Em Structure Of Candida Albicans Vrg4 Bound To An Inhibitory Nanobody And Gdp-Mannose.
Organism: Candida albicans, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-09-03 Classification: TRANSPORT PROTEIN Ligands: GDD |

