Search Count: 90
All
Selected
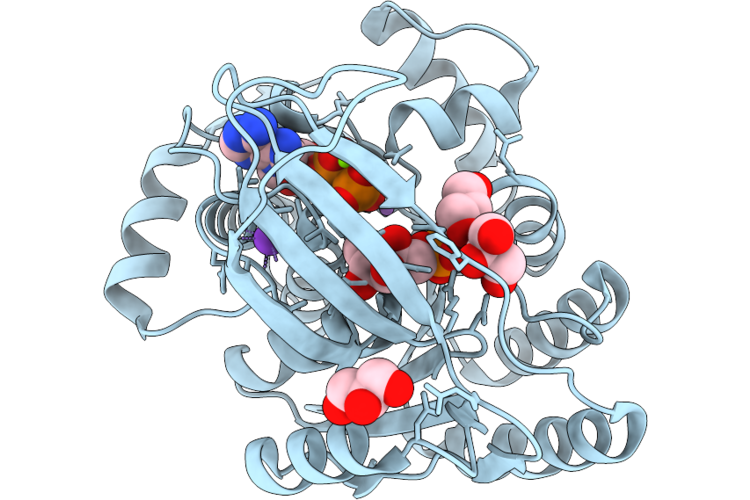 |
1-Phosphofructokinase (Fruk) From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, F1X, ADP, LI, K, GOL |
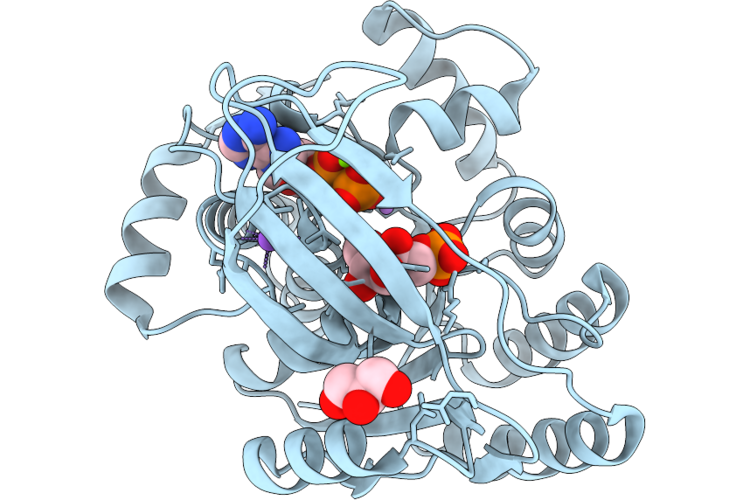 |
1-Phosphofructokinase Mutant K95H From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, LI, NA, ADP, F1X, GOL |
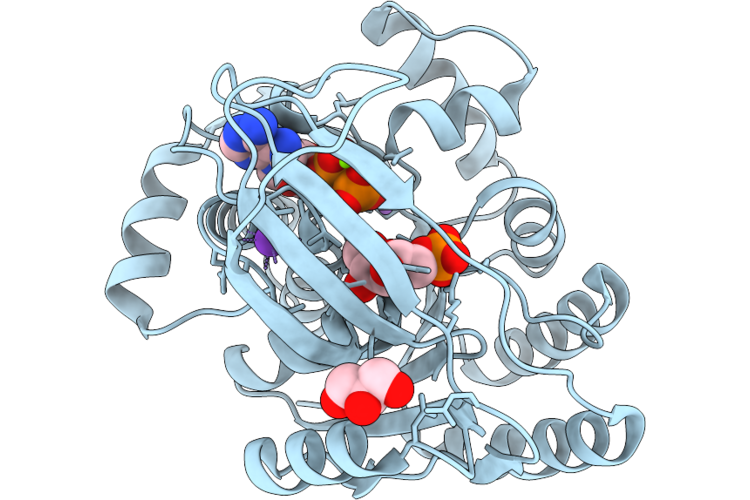 |
1-Phosphofructokinase Mutant K95H From E. Coli With Bound Fructose 6-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GOL, ADP, F6P, MG, K, LI |
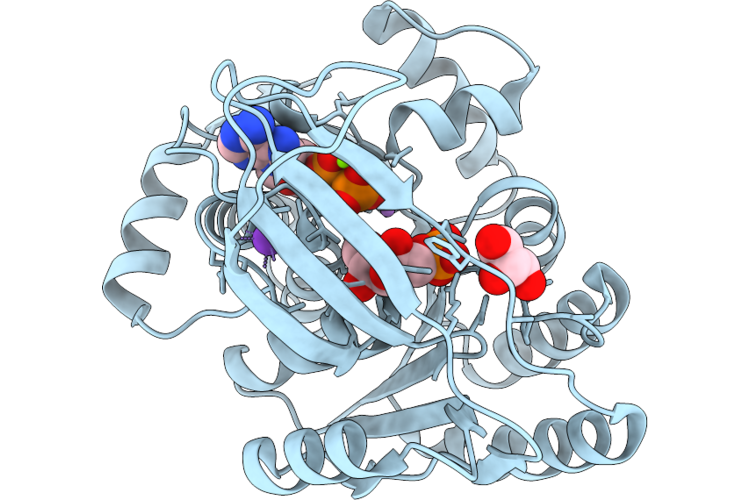 |
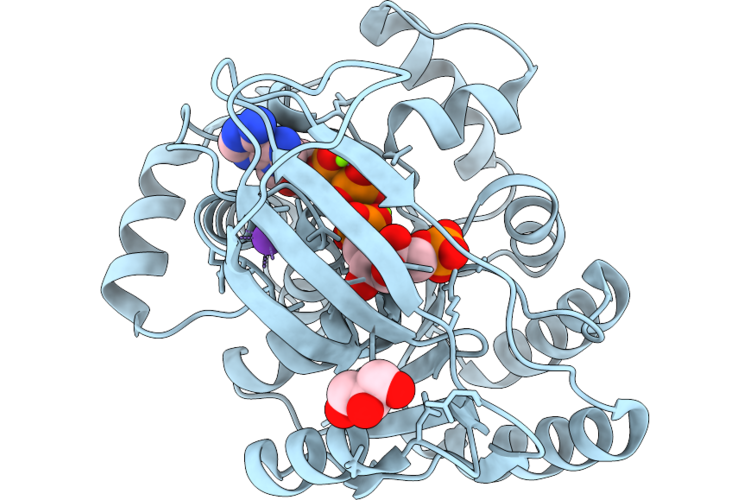
1-Phosphofructokinase Mutant K95T/G110S From E. Coli With Bound Fructose 1-Phosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, F1X, ADP, K, LI, GOL |
 |
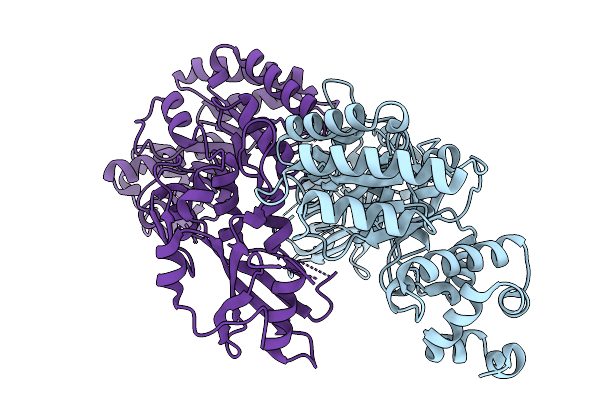
1-Phosphofructokinase Mutant K95T/G110S From E. Coli With Bound Fructose-1,6-Bisphosphate And Adp
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: MG, ADP, FBP, GOL, K |
 |
Organism: Rhodospirillum rubrum atcc 11170
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-02-04 Classification: LYASE |
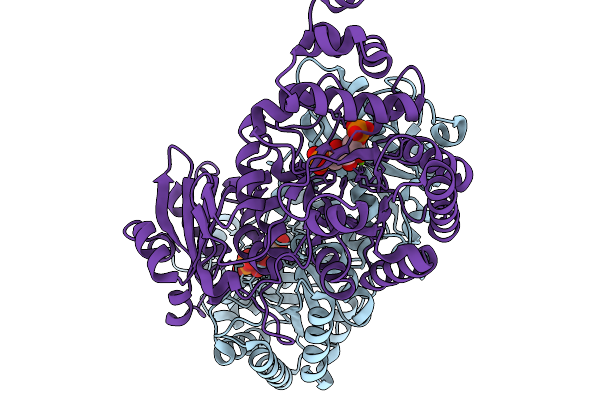 |
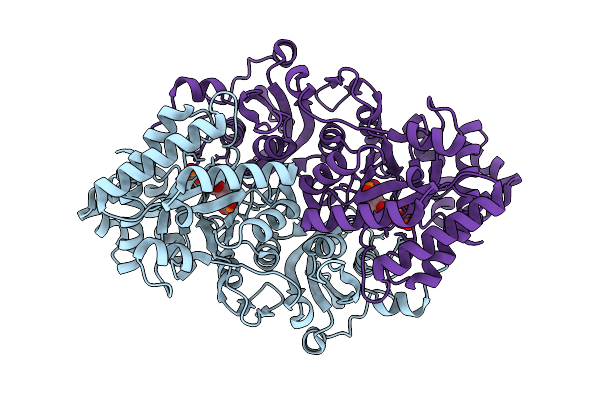
Activated Form Ii Rubisco From Rhodospirillum Rubrum With Bound Magnesium And Cabp
Organism: Rhodospirillum rubrum atcc 11170
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-02-04 Classification: LYASE Ligands: MG, CAP |
 |
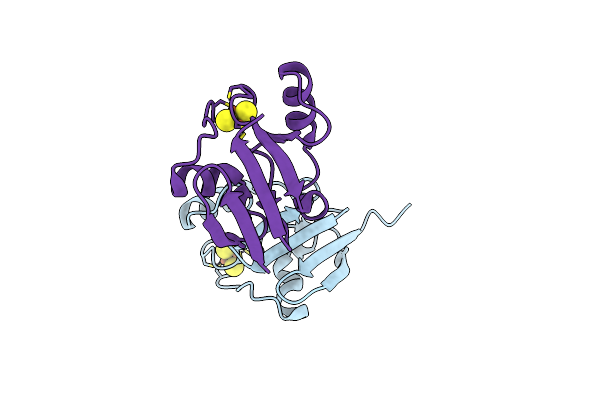
Form Ii Rubisco Inside Epyc1-Formed Liquid-Liquid Phase Separated Condensates With Bound Magnesium And Cabp
Organism: Rhodospirillum rubrum atcc 11170
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: LYASE Ligands: MG, CAP |
 |
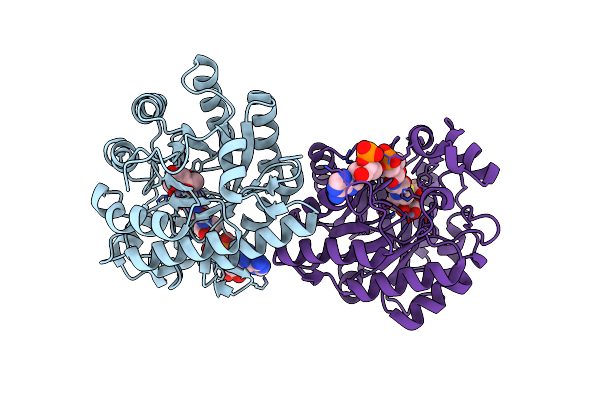
Organism: Rhodobacter capsulatus sb 1003
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-06-04 Classification: ELECTRON TRANSPORT Ligands: FES |
 |
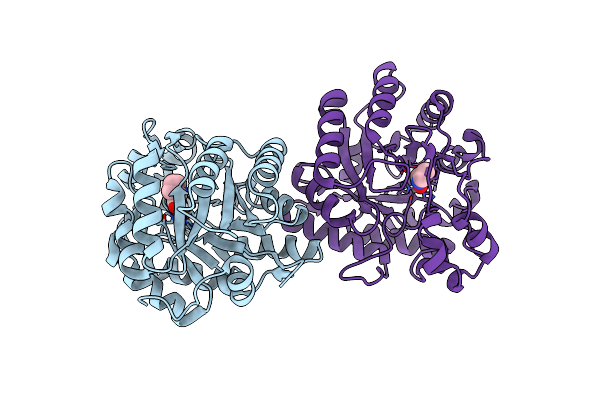
Beta-Keto Acid Cleavage Enzyme From Paracoccus Denitrificans With Bound Acetoacetate And Acetyl-Coa
Organism: Paracoccus denitrificans pd1222
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-04-16 Classification: LYASE Ligands: ACO, AAE, ZN |
 |
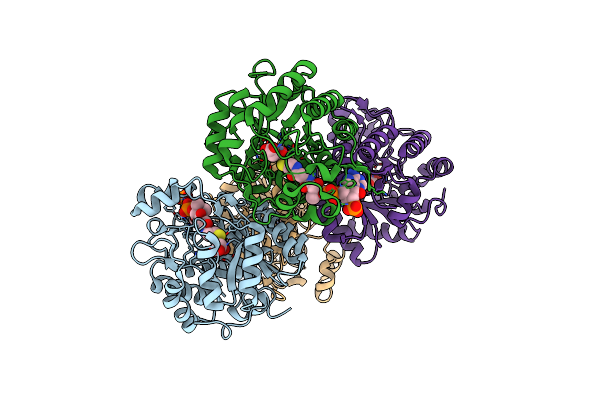
Organism: Paracoccus denitrificans pd1222
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-01-01 Classification: LYASE Ligands: ZN, PRO |
 |
Beta-Keto Acid Cleavage Enzyme From Paracoccus Denitrificans With Bound Malonate And Coenzyme A
Organism: Paracoccus denitrificans pd1222
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2025-01-01 Classification: LYASE Ligands: MLI, COA, ZN |
 |
L2 Forming Rubisco Derived From Ancestral Sequence Reconstruction Of The Last Common Ancestor Of Form I'' And Form I Rubiscos
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-10-16 Classification: LYASE Ligands: CAP, MG |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-10-02 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
Succinic Semialdehyde Dehydrogenase From E. Coli With Bound Nad+ And Succinic Semialdehyde
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-10-02 Classification: OXIDOREDUCTASE Ligands: NAD, SSN |
 |
Succinic Semialdehyde Dehydrogenase From E. Coli With Q262R Substitution And Bound Nad+
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-10-02 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
Succinic Semialdehyde Dehydrogenase From E. Coli With Q262R Substitution And Bound Nad+, Succinic Semialdehyde
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-10-02 Classification: OXIDOREDUCTASE Ligands: NAD, SSN |
 |
Non-Obligately L8S8-Complex Forming Rubisco Derived From Ancestral Sequence Reconstruction And Rational Engineering In L8S8 Complex With Substitutions R269W, E271R, L273N
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-10-02 Classification: LYASE Ligands: CAP, MG, B3P |
 |
Crystal Structure Of 2-Hydroxacyl-Coa Lyase/Synthase Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, L-Lactyl-Coa, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2024-10-02 Classification: LYASE Ligands: 8FL, UQ3, ADP, ACE, MG, GOL |
 |
Crystal Structure Of 2-Hydroxacyl-Coa Lyase/Synthase Apbhacs From Alphaproteobacteria Bacterium In The Complex With Thdp, D-Lactyl-Coa, And Adp
Organism: Alphaproteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2024-10-02 Classification: LYASE Ligands: MG, 8FL, UQ3, ADP, EDO, ACY, PO4, CL, ACE, PEG |

