Search Count: 221
All
Selected
 |
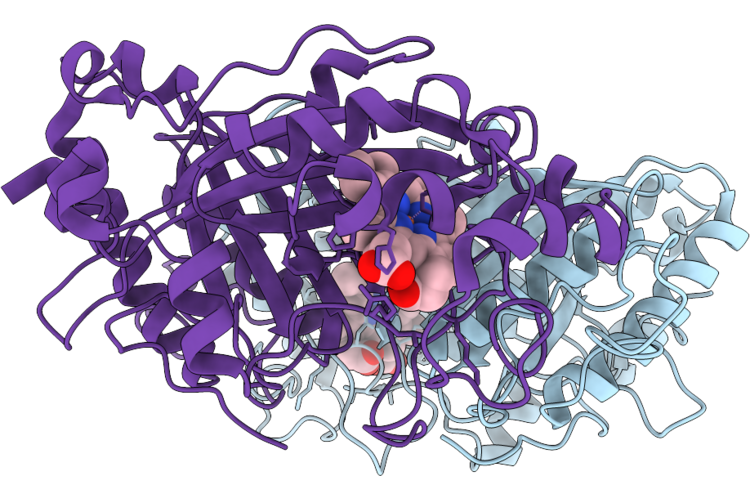
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
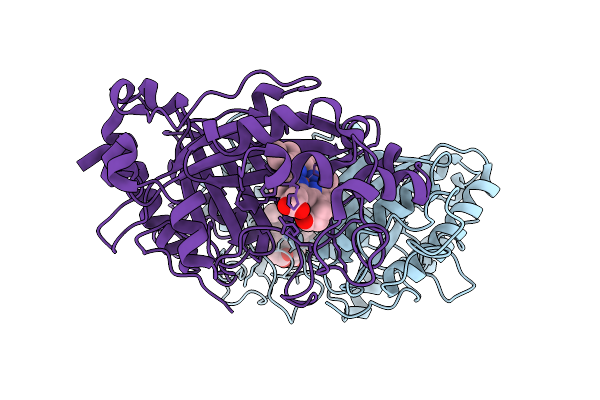
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2025-10-29 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
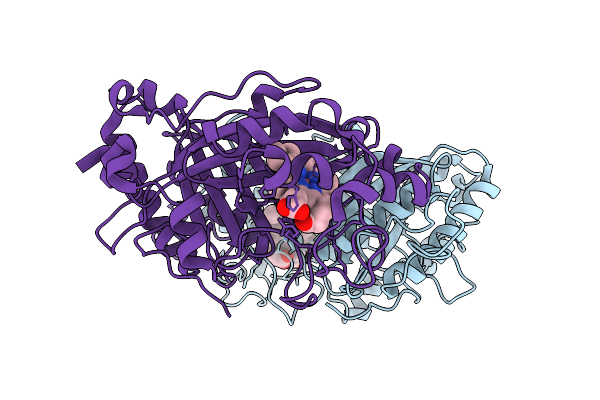
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
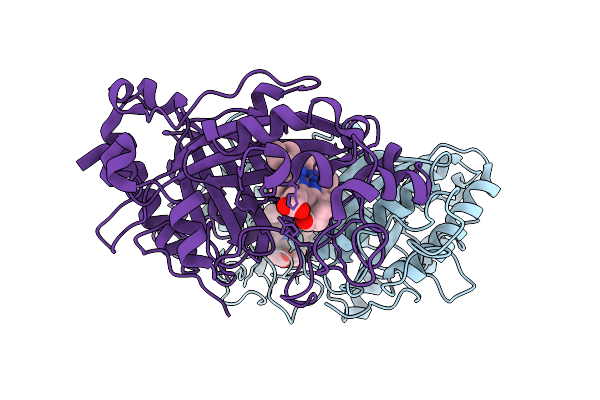
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
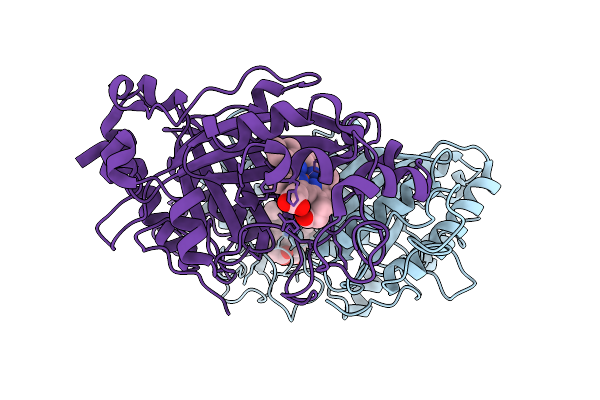 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: HEM |
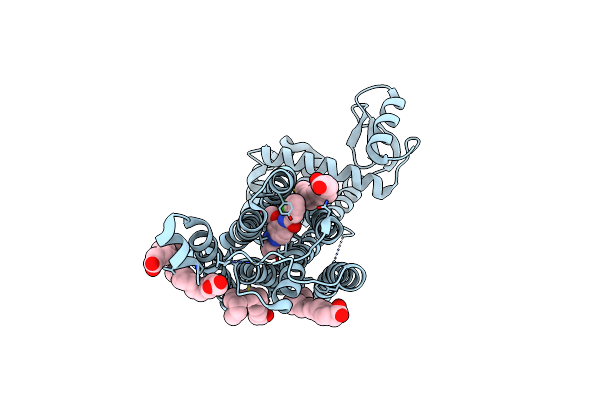 |
Dark Structure Of The Human Metabotropic Glutamate Receptor 5 Transmembrane Domain Bound To Photoswitchable Ligand Alloswitch-1
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2025-06-25 Classification: SIGNALING PROTEIN Ligands: OLA, 4YI |
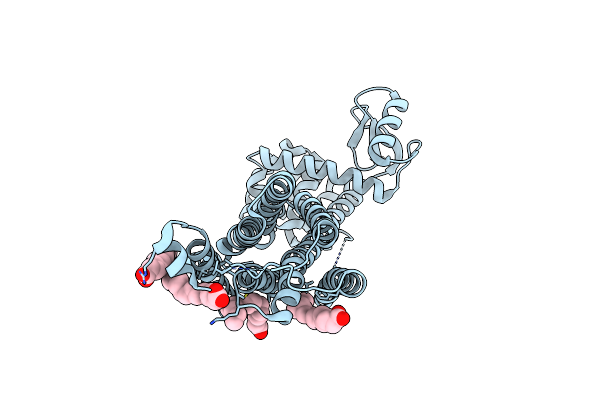 |
Apo-State Structure Of The Human Metabotropic Glutamate Receptor 5 Transmembrane Domain Freeze-Trapped After Light Activation Of Photoswitchable Ligand Alloswitch-1
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2025-06-25 Classification: SIGNALING PROTEIN Ligands: OLA |
 |
Grouped 150-240 Ms Dark Structure Of Sensory Rhodopsin-Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL, BOG, MPG |
 |
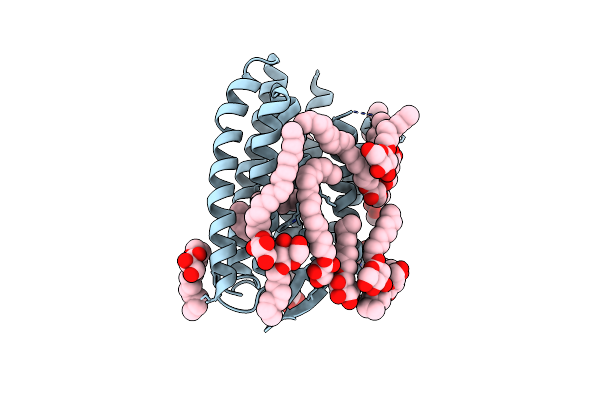
Continuously Illuminated Structure Of Sensory Rhodopsin Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL |
 |
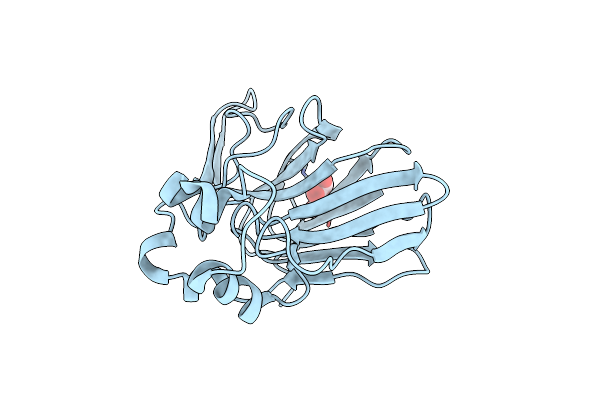
Continuous Dark State Structure Of Sensory Rhodopsin Ii Solved By Serial Millisecond Crystallography
Organism: Natronomonas pharaonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-04-30 Classification: SIGNALING PROTEIN Ligands: RET, CL, BOG, MPG |
 |
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-04-02 Classification: PLANT PROTEIN Ligands: TLA |
 |
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:1.46 Å Release Date: 2025-04-02 Classification: PLANT PROTEIN Ligands: TLA |
 |
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-04-02 Classification: PLANT PROTEIN Ligands: TLA |
 |
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2025-04-02 Classification: PLANT PROTEIN Ligands: TLA |

