Search Count: 14
All
Selected
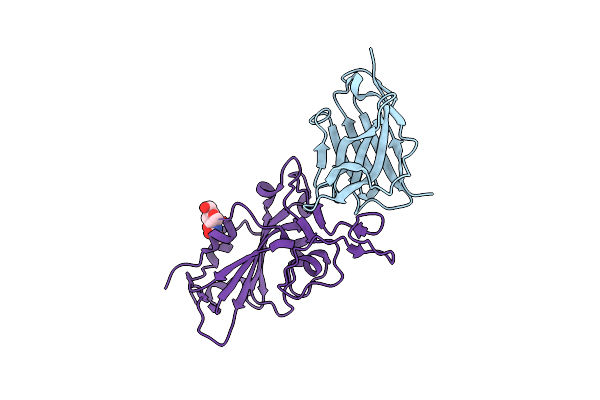 |
Organism: Vicugna pacos, Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-08-10 Classification: VIRAL PROTEIN/ANTIVIRAL PROTEIN Ligands: NAG |
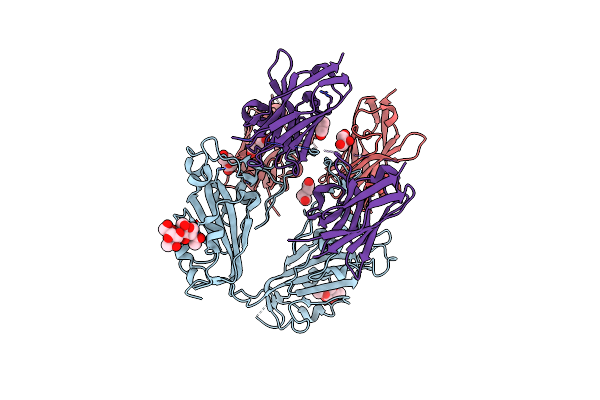 |
Crystallographic Structure Of Two Neutralizing Nanobodies In Complex With Sars-Cov-2 Spike Receptor-Binding Domain (Rbd)
Organism: Severe acute respiratory syndrome coronavirus 2, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2022-08-03 Classification: ANTIVIRAL PROTEIN Ligands: NAG, PEG, GOL |
 |
Crystal Structure Of The Sars-Cov-2 (Covid-19) Main Protease In Complex With Uaw243
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2020-06-24 Classification: HYDROLASE Ligands: GOL |
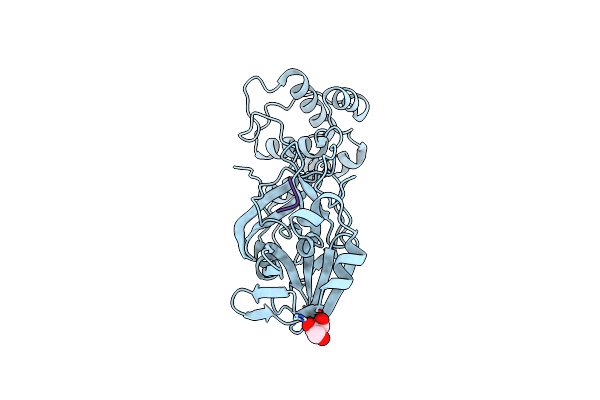 |
Crystal Structure Of The Sars-Cov-2 (Covid-19) Main Protease In Complex With Uaw241
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2020-06-17 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: GOL |
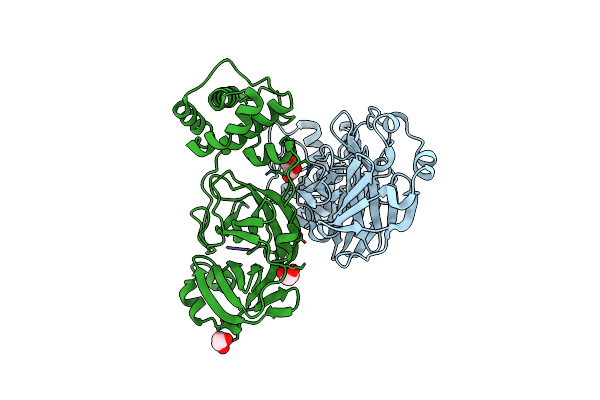 |
Crystal Structure Of The Sars-Cov-2 (Covid-19) Main Protease In Complex With Inhibitor Uaw246
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2020-06-17 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: GOL, NA |
 |
Crystal Structure Of The Sars-Cov-2 (Covid-19) Main Protease In Complex With Inhibitor Uaw247
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2020-06-17 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: GOL, NA |
 |
Crystal Structure Of The Sars-Cov-2 (Covid-19) Main Protease In Complex With Inhibitor Uaw248
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2020-06-17 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: GOL, NA, DMS |
 |
Model Of Human Anaphase-Promoting Complex/Cyclosome (Apc/C-Cdh1) With E2 Ube2S Poised For Polyubiquitination Where Ube2S, Apc2, And Apc11 Are Modeled Into Low Resolution Density
Organism: Homo sapiens, Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2016-10-26 Classification: CELL CYCLE |
 |
Model Of Human Anaphase-Promoting Complex/Cyclosome (Apc/C-Cdh1) With A Cross Linked Ubiquitin Variant-Substrate-Ube2C (Ubch10) Complex Representing Key Features Of Multiubiquitination
Organism: Homo sapiens, Saccharomyces cerevisiae, Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2016-09-14 Classification: SIGNALING PROTEIN |
 |
Model Of Human Anaphase-Promoting Complex/Cyclosome (Apc15 Deletion Mutant), In Complex With The Mitotic Checkpoint Complex (Apc/C-Cdc20-Mcc) Based On Cryo Em Data At 4.8 Angstrom Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2016-09-07 Classification: CELL CYCLE |
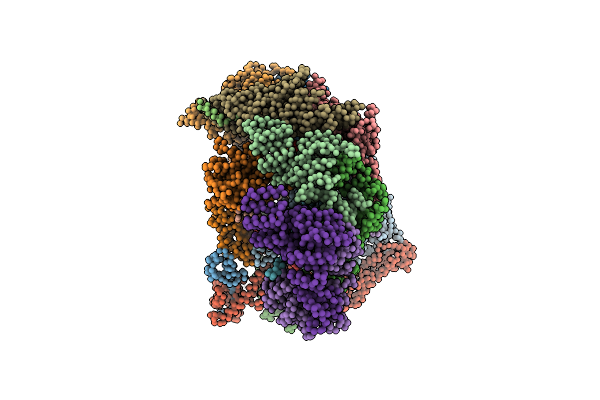 |
Model Of Human Anaphase-Promoting Complex/Cyclosome Complex (Apc15 Deletion Mutant) In Complex With The E2 Ube2C/Ubch10 Poised For Ubiquitin Ligation To Substrate (Apc/C-Cdc20-Substrate-Ube2C)
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2016-08-24 Classification: CELL CYCLE |
 |
Apc11-Ubv Shows Role Of Noncovalent Ring-Ubiquitin Interactions In Processive Multiubiquitination And Ubiquitin Chain Elongation By Apc/C
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2016-06-15 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2014-10-29 Classification: METAL BINDING PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2014-10-29 Classification: LIGASE Ligands: ZN |

