Search Count: 49
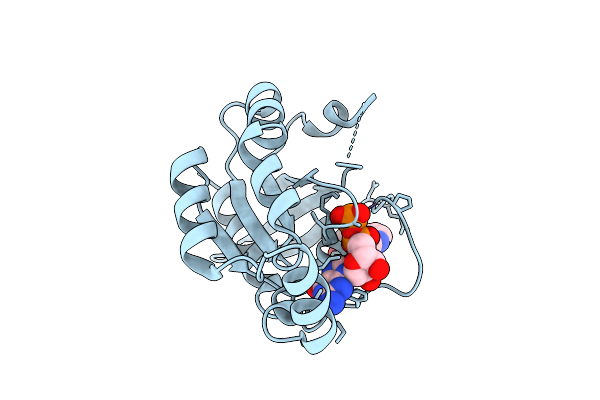 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, A1D9L, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
 |
Structure Of The Rhoa Y42C Mutant Featuring Thr37 Coordination Of Magnesium Ion
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-06-04 Classification: CELL INVASION Ligands: GDP, MG |
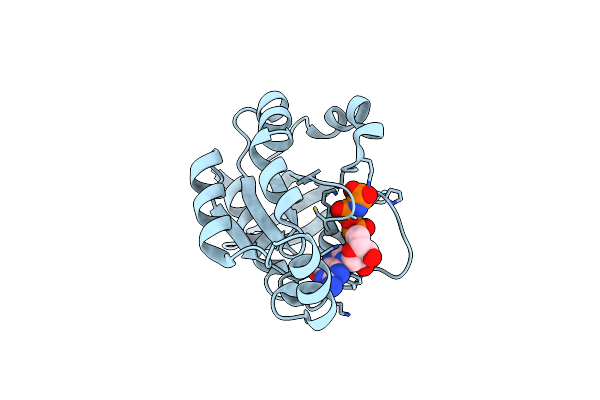 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-06-04 Classification: CELL INVASION Ligands: GNP, MG |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-04-30 Classification: STRUCTURAL PROTEIN |
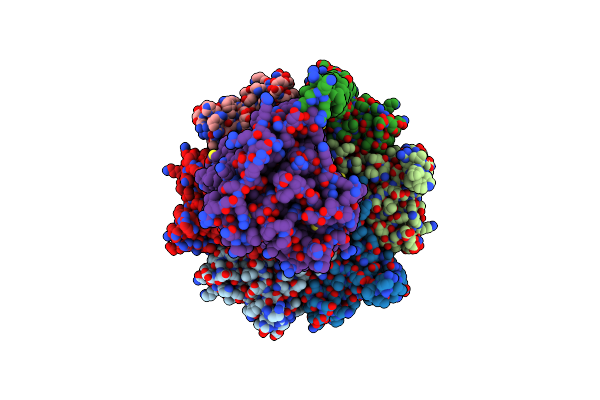 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-04-30 Classification: STRUCTURAL PROTEIN |
 |
Organism: Autographa californica nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Organism: Autographa californica nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: NAG |
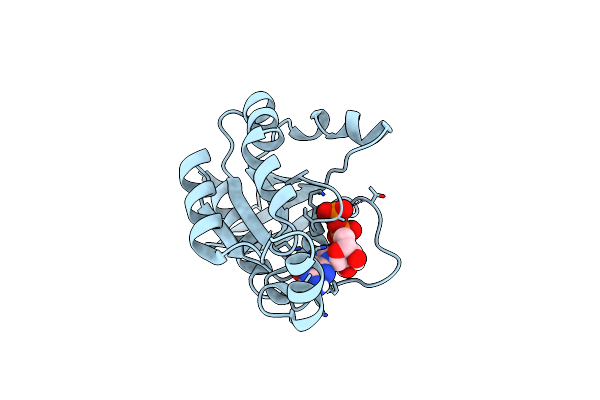 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2023-02-22 Classification: CELL INVASION Ligands: GDP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-02-22 Classification: CELL INVASION Ligands: GOL, GDP, MG |
 |
Organism: Penicillium rubens wisconsin 54-1255
Method: SOLUTION NMR Release Date: 2022-08-24 Classification: ANTIMICROBIAL PROTEIN |
 |
Organism: Penicillium rubens (strain atcc 28089 / dsm 1075 / nrrl 1951 / wisconsin 54-1255)
Method: SOLUTION NMR Release Date: 2022-03-30 Classification: ANTIMICROBIAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2022-02-16 Classification: ISOMERASE Ligands: J2C, PE8 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2022-02-16 Classification: ISOMERASE Ligands: J3X |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-02-16 Classification: ISOMERASE Ligands: J50, PE8 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-02-16 Classification: ISOMERASE Ligands: 0BF, P33 |
 |
Cryo-Em Structure Of Pdr5 From Saccharomyces Cerevisiae In Inward-Facing Conformation Without Nucleotides
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2021-11-10 Classification: TRANSPORT PROTEIN |
 |
Cryo-Em Structure Of Pdr5 From Saccharomyces Cerevisiae In Inward-Facing Conformation With Adp/Atp
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2021-11-10 Classification: TRANSPORT PROTEIN Ligands: ATP, ADP |
 |
Cryo-Em Structure Of Pdr5 From Saccharomyces Cerevisiae In Inward-Facing Conformation With Adp/Atp And Rhodamine 6G
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2021-11-10 Classification: TRANSPORT PROTEIN Ligands: ATP, ADP, RHQ |

