Search Count: 174
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
 |
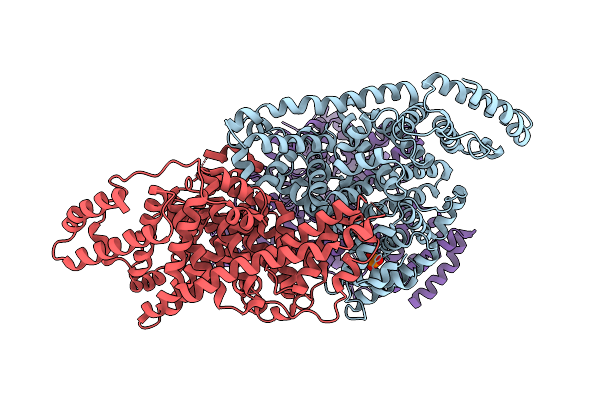
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
 |
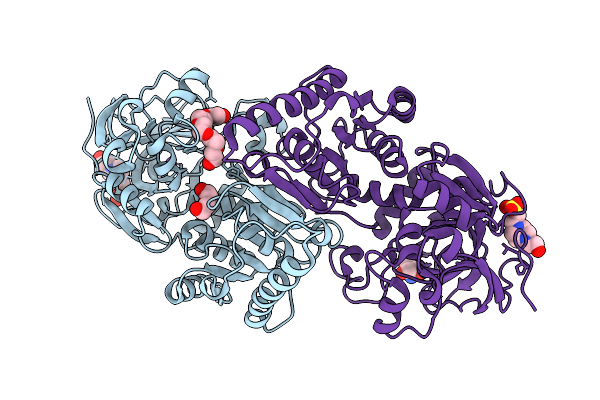
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-01-21 Classification: TRANSPORT PROTEIN Ligands: PO4 |
 |
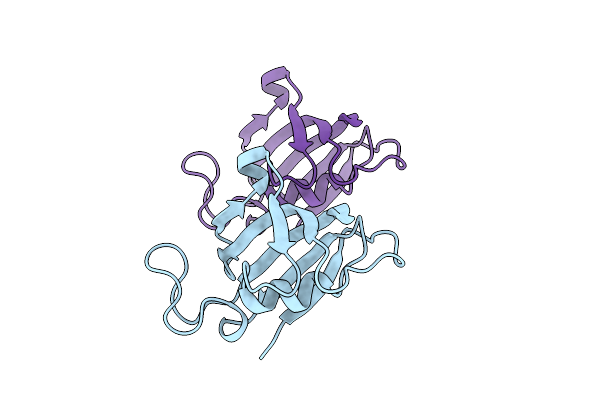
Organism: Mesorhizobium metallidurans
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: P6G, PEG, EPE, NA, TRS |
 |
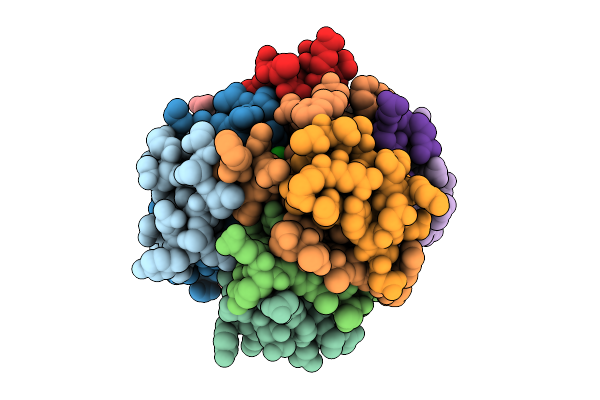
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-11-05 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-10-29 Classification: HORMONE Ligands: CRS, ZN, CL, CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2025-10-29 Classification: HORMONE Ligands: IPH, ZN, CL, CA |
 |
3-Hydroxypropionyl-Coa Synthetase (Adp-Forming) From Nitrosopumilus Maritimus.
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-10-01 Classification: LIGASE Ligands: ANP, 3OH, PO4, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-09-03 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-09-03 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-09-03 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
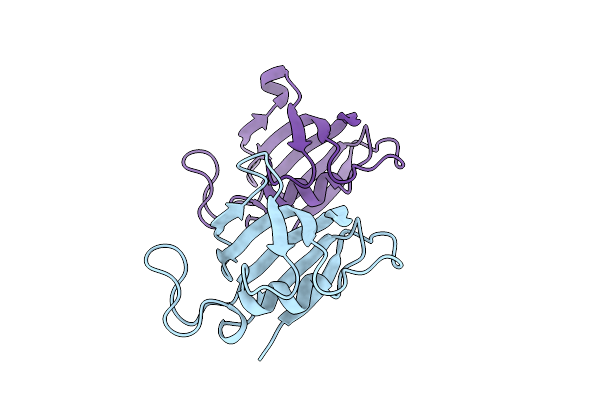 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
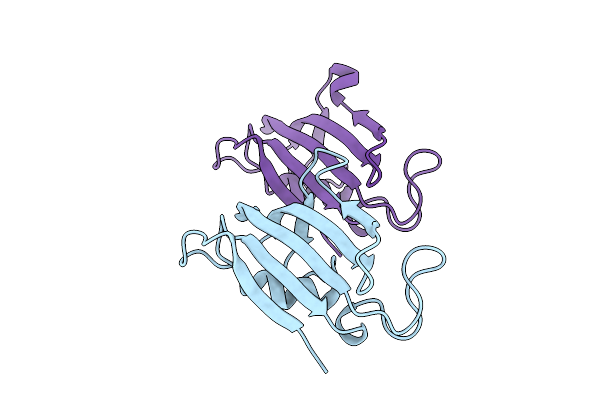 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
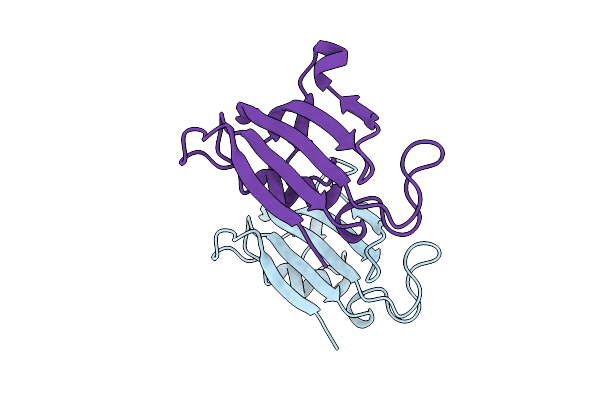 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: ISOMERASE |
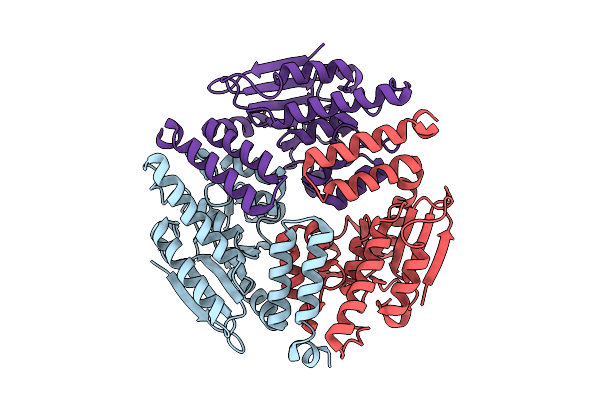 |
Cryogenic Temperature Crystal Structure Of Nmar_1308 Protein At 2.46 Angstrom Resolution
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2025-07-09 Classification: STRUCTURAL PROTEIN |

