Search Count: 949
All
Selected
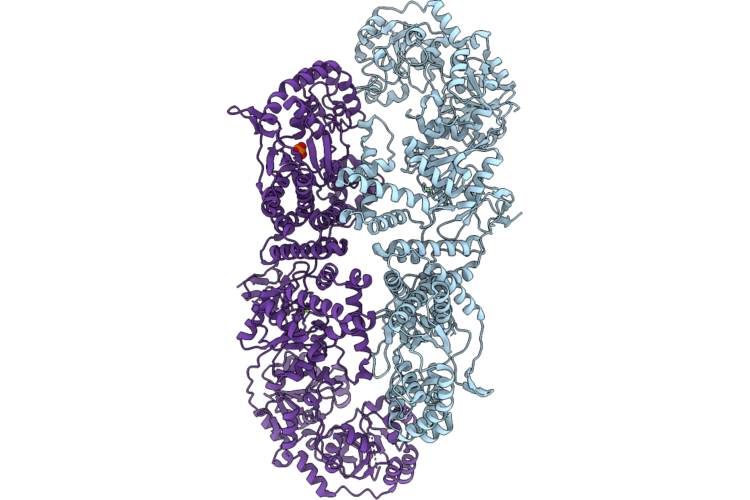 |
Organism: Haemophilus influenzae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: MG, PO4 |
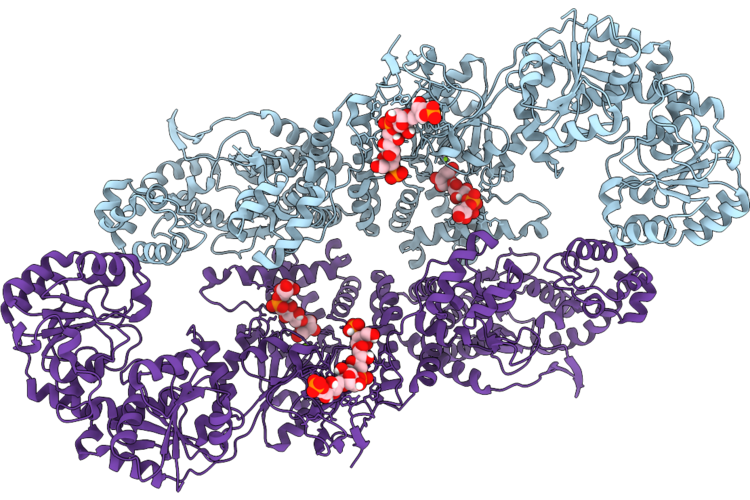 |
Haemophilus Influenzae Serotype B Capsule Polymerase Bcs3 In Complex With Acceptor Polymer
Organism: Haemophilus influenzae
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: BIOSYNTHETIC PROTEIN Ligands: MG, KOF |
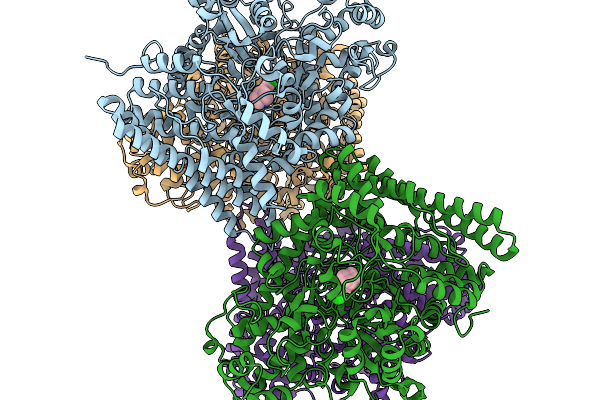 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: A1JCI |
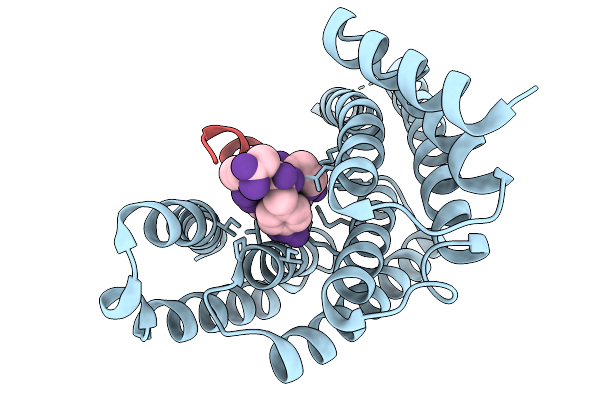 |
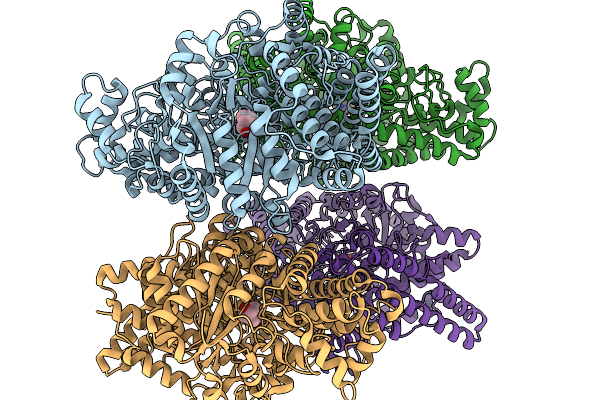
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: PROTEIN BINDING |
 |
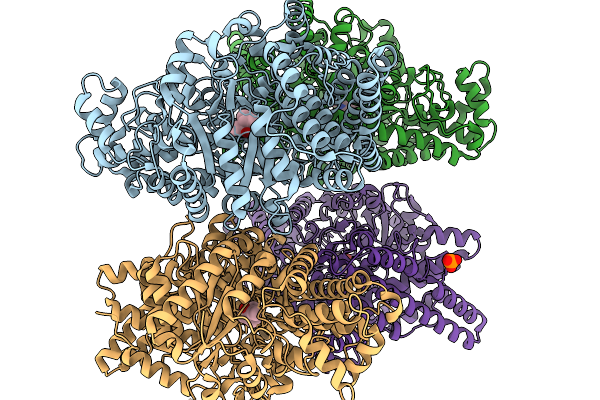
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXC |
 |
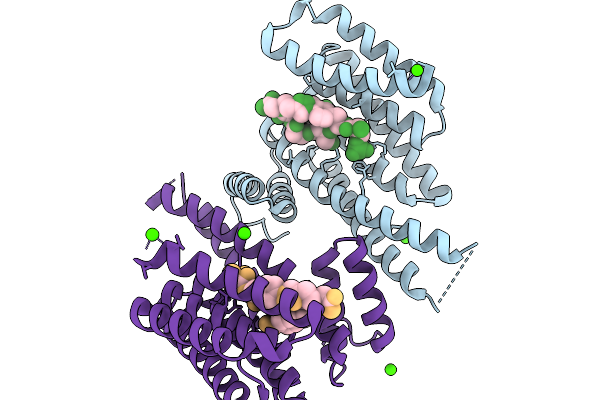
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXB, PO4 |
 |
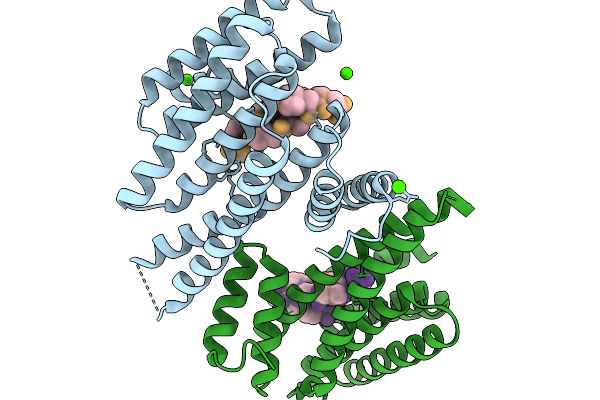
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-01-14 Classification: PROTEIN BINDING Ligands: CA, CL |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-01-14 Classification: PROTEIN BINDING Ligands: CL, CA |
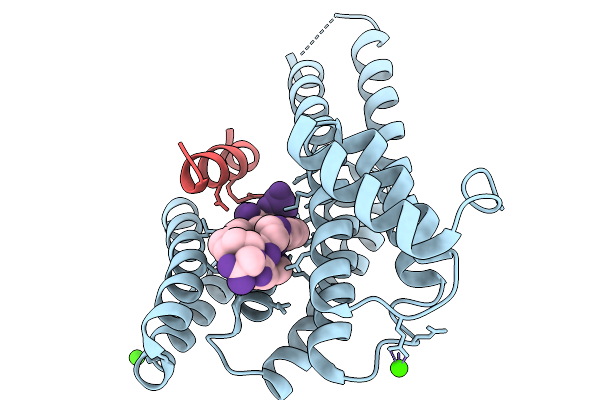 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-01-14 Classification: PROTEIN BINDING Ligands: CA |
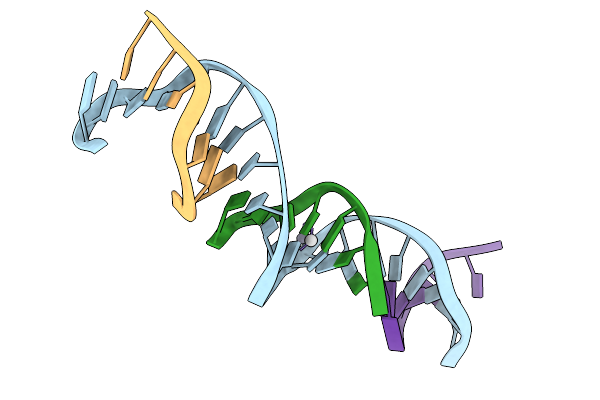 |
[T:Ag+/Hg2+:T--(Ph8-Ph9.5; 50S)] Metal-Mediated Dna Base Pair In Tensegrity Triangle Grown At Ph 8 And Soaked In Ph 9.5 Ag/Hg For 50S
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:4.89 Å Release Date: 2025-12-31 Classification: DNA Ligands: HG, AG |
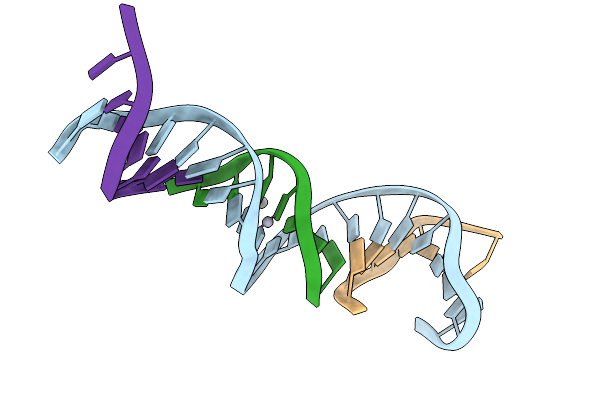 |
[T:Ag+/Hg2+:T--(Ph8-Ph9.5; 75S)] Metal-Mediated Dna Base Pair In Tensegrity Triangle Grown At Ph 8 And Soaked In Ph 9.5 For 75S
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:5.70 Å Release Date: 2025-12-31 Classification: DNA Ligands: HG, AG |
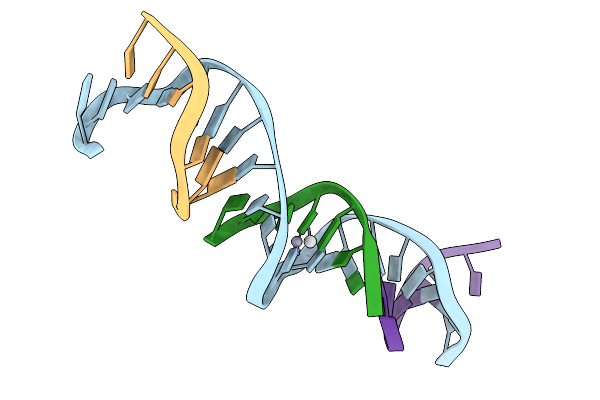 |
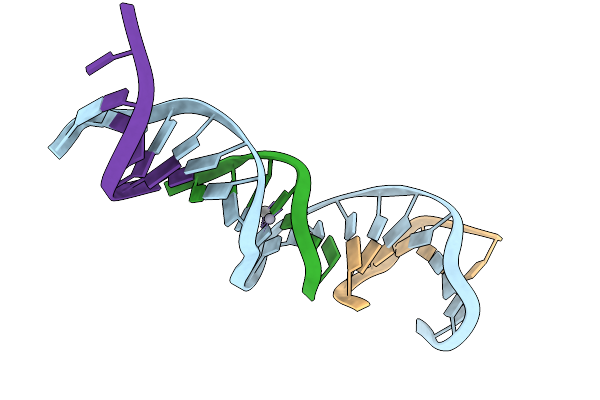
[T:Ag+/Hg2+:T--(Ph8-Ph9.5; 95S)] Metal-Mediated Dna Base Pair In Tensegrity Triangle Grown At Ph 8 And Soaked In Ph 9.5 Ag/Hg For 95S
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:5.38 Å Release Date: 2025-12-31 Classification: DNA Ligands: HG, AG |
 |
[T:Ag+/Hg2+:T--(Ph11-Ph9.5; 5S)] Metal-Mediated Dna Base Pair In Tensegrity Triangle Grown At Ph 8 And Soaked In Ph 9.5 Ag/Hg For 5S
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:4.66 Å Release Date: 2025-12-31 Classification: DNA Ligands: AG, HG |
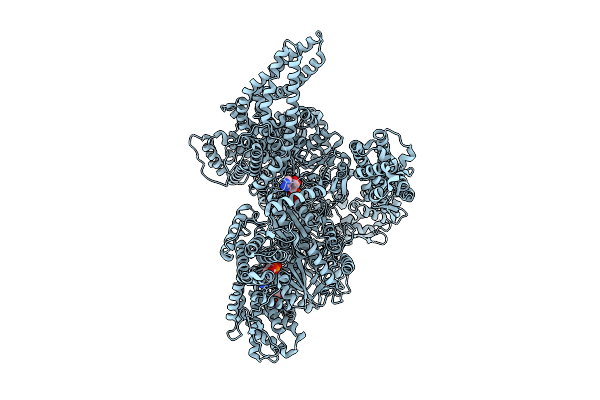 |
Motor Domain With Apo Aaa1 And Adp Aaa3 From Yeast Full-Length Dynein-1 In 0.1 Mm Atp Condition
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MOTOR PROTEIN Ligands: ATP, ADP, MG |
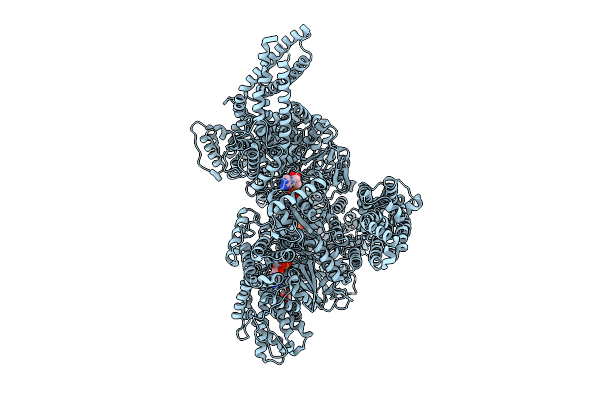 |
Motor Domain With Adp Aaa1 And Adp Aaa3 From Yeast Full-Length Dynein-1 In 0.1 Mm Atp Condition
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MOTOR PROTEIN Ligands: ADP, ATP |
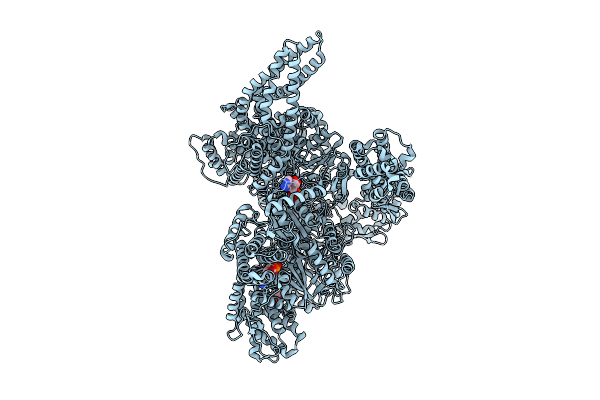 |
Motor Domain Alone With Apo Aaa1 And Adp Aaa3 From Yeast Full-Length Dynein-1 And Pac1 In 0.1 Mm Atp Condition
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-12-17 Classification: MOTOR PROTEIN Ligands: ATP, ADP, MG |
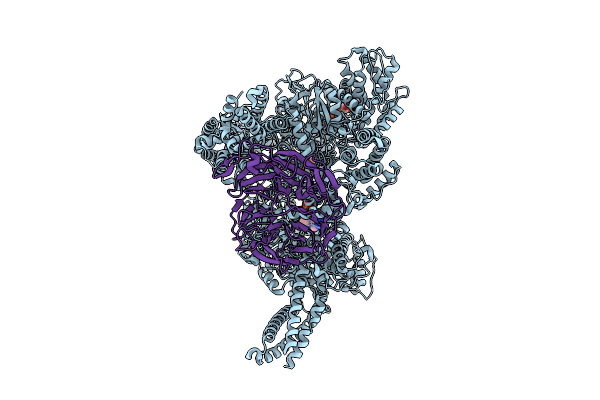 |
Motor Domain-Pac1 Complex With Adp Aaa1 And Apo Aaa3 From Yeast Full-Length Dynein-1 And Pac1 In 0.1 Mm Atp Condition
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-12-17 Classification: MOTOR PROTEIN Ligands: ADP, ATP, MG |
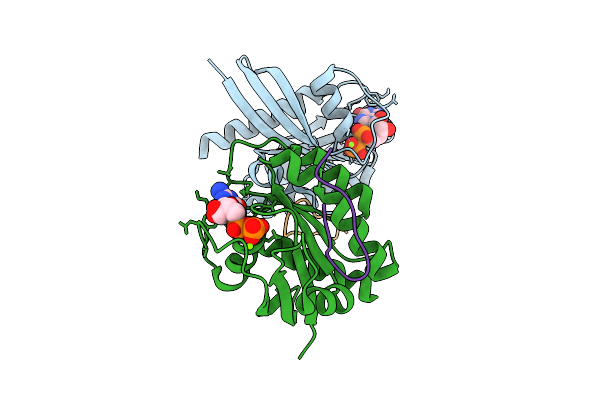 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: MG, GDP |
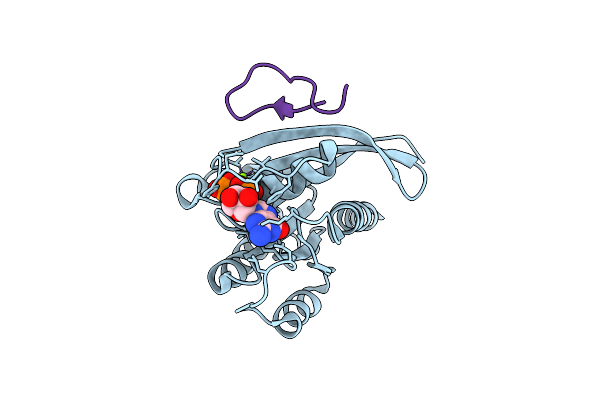 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: MG, GTP |
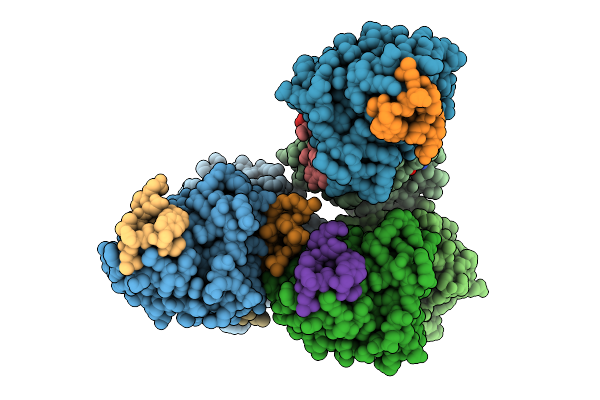 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-10 Classification: HYDROLASE Ligands: MG, GTP |

