Search Count: 8
All
Selected
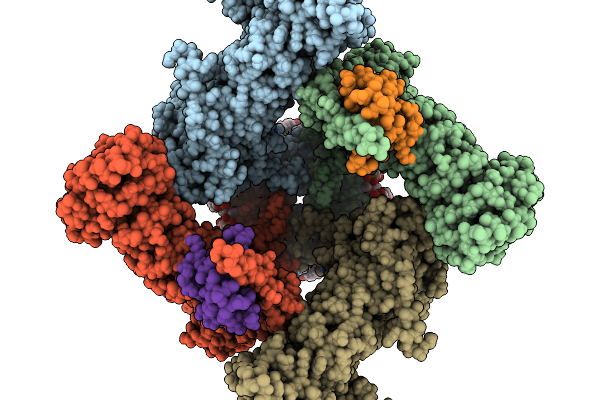 |
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, CA, D39 |
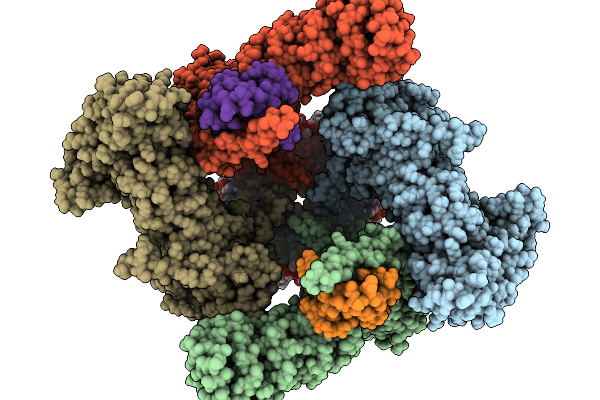 |
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, CA, D39 |
 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Nicotinamide, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, NCA, D39 |
 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen And Calcium
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, CA, D39 |
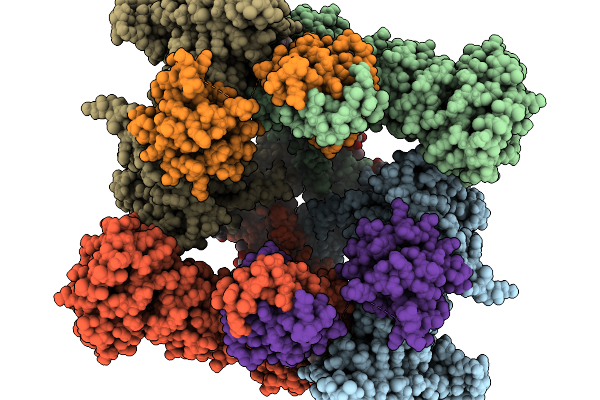 |
Structure Of Nanchung-Inactive-Calmodulin In Complex With Afidopyropen, Edta
Organism: Halyomorpha halys, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-02-11 Classification: TRANSPORT PROTEIN Ligands: 6OU, LBN, A1B32, D39 |
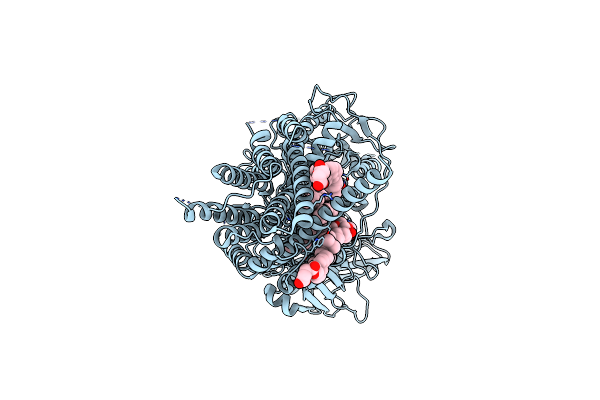 |
Cryo-Em Structure Of The Gpi Inositol-Deacylase (Pgap1/Bst1) From Chaetomium Thermophilum
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719), Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-12-20 Classification: MEMBRANE PROTEIN Ligands: Y01, D39 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: MEMBRANE PROTEIN Ligands: VN4, MG, D39, NAG |
 |
Cryo-Em Structure Of The Human P4-Type Flippase Atp8A1-Cdc50 (E2Pi-Pl State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2019-08-28 Classification: MEMBRANE PROTEIN Ligands: ALF, Y01, MG, D39, NAG |

