Search Count: 142
All
Selected
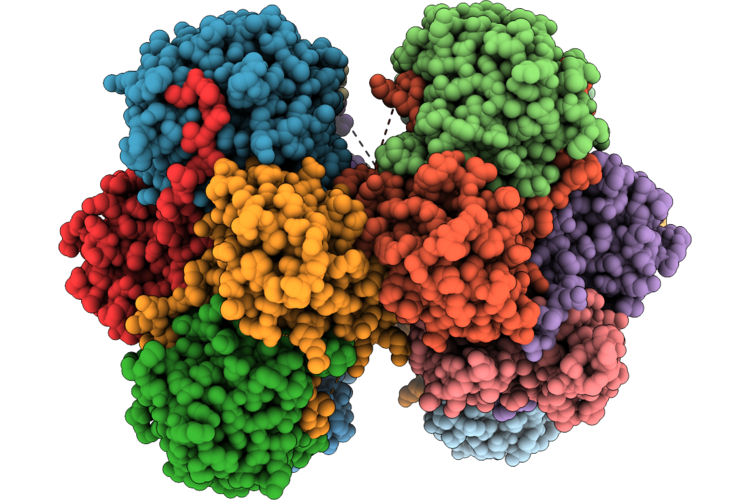 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-04-22 Classification: HYDROLASE |
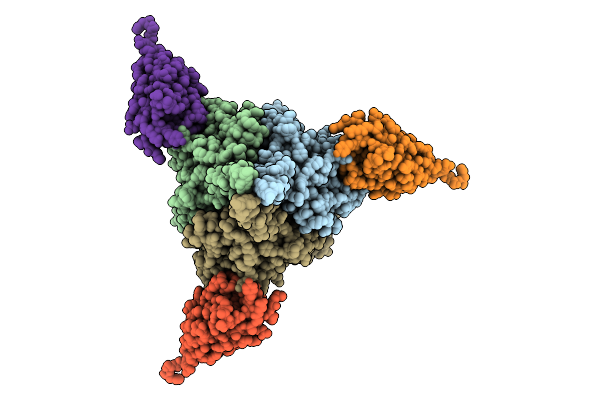 |
Cryo-Em Structure Of Prefusion-Stabilized Rsv F (Ds-Cav1 Strain: A2) In Complex With Nanobody 1G9
Organism: Respiratory syncytial virus a2, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: ANTIVIRAL PROTEIN/IMMUNE SYSTEM |
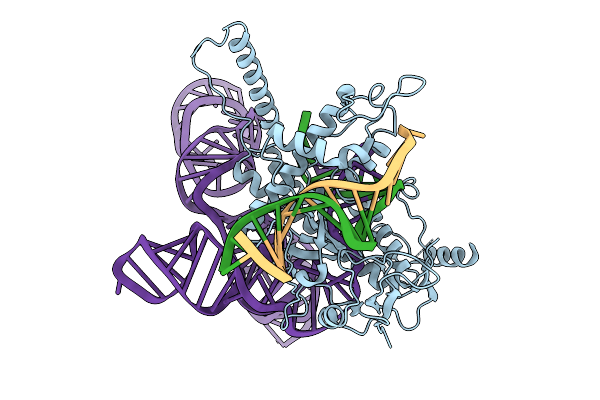 |
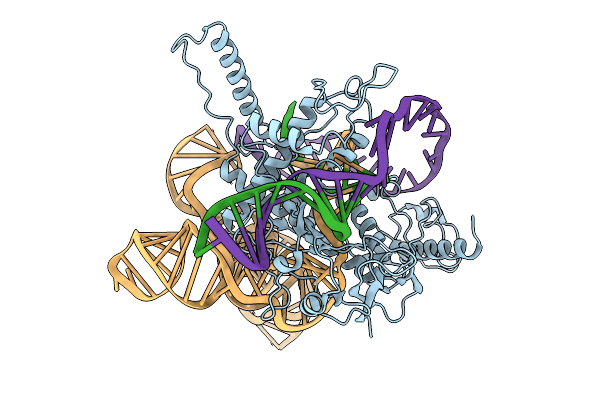
Structure Of Latranc Complex Bound To 27Nt Complementary Dna Substrate, Conformation 1
Organism: Lawsonibacter sp.
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: IMMUNE SYSTEM/DNA/RNA |
 |
Organism: Lawsonibacter sp.
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: IMMUNE SYSTEM/DNA/RNA |
 |
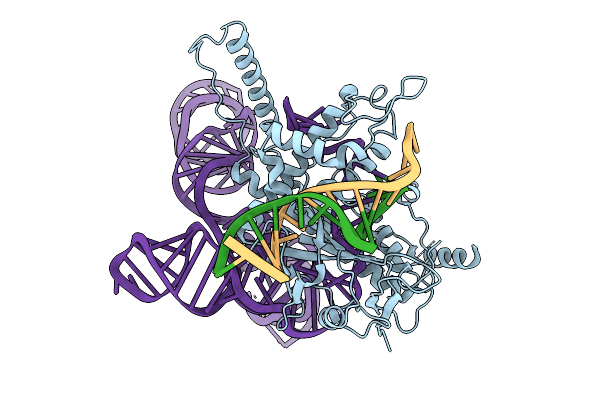
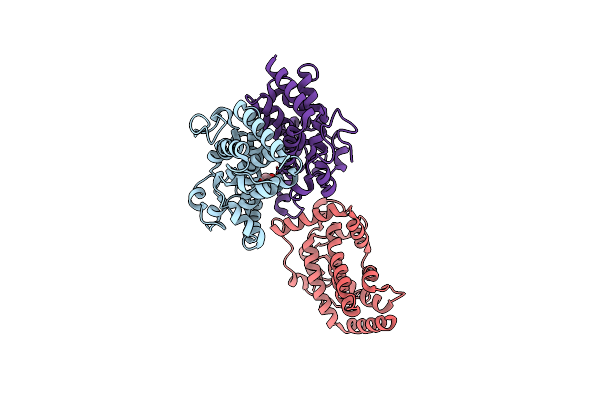
Structure Of Latranc Complex Bound To 27Nt Complementary Dna Substrate, Conformation 2
Organism: Lawsonibacter sp.
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: IMMUNE SYSTEM/DNA/RNA |
 |
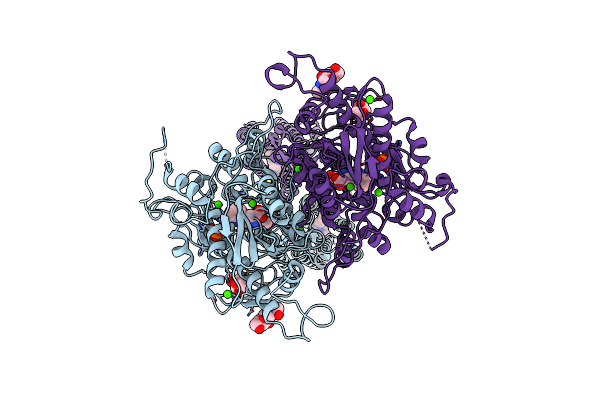
Organism: Shigella flexneri
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-02-19 Classification: HYDROLASE Ligands: GOL |
 |
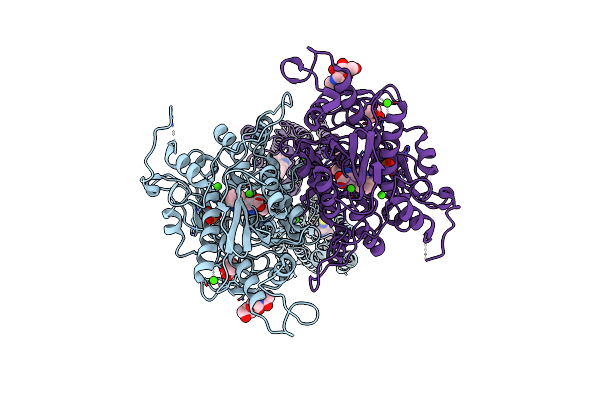
Structure Of Calcium-Sensing Receptor In Complex With Positive Allosteric Modulator '6218
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-10-02 Classification: MEMBRANE PROTEIN Ligands: NAG, CA, TRP, PO4, A1ATP |
 |
Structure Of Calcium-Sensing Receptor In Complex With Positive Allosteric Modulator '54149
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-10-02 Classification: MEMBRANE PROTEIN Ligands: NAG, CA, TRP, PO4, A1ATX |
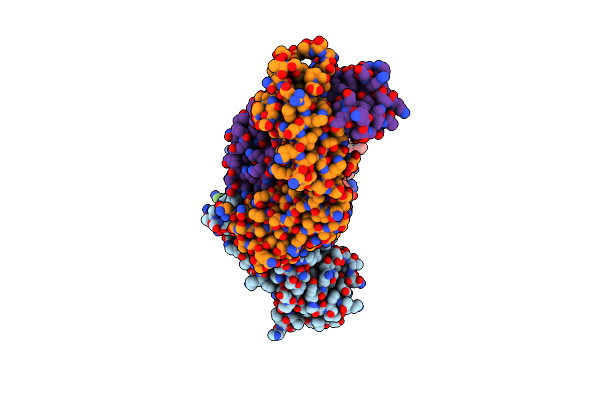 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
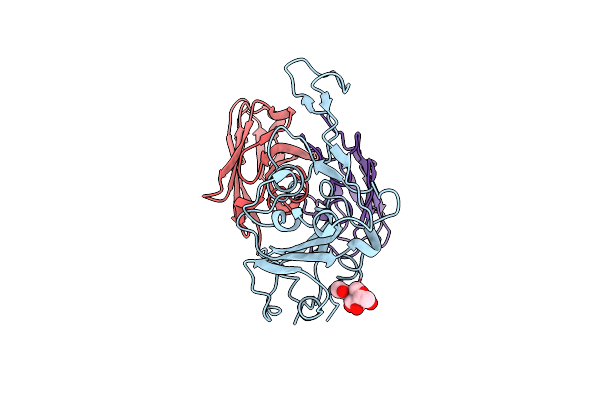
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: VIRAL PROTEIN/HYDROLASE/IMMUNE SYSTEM Ligands: NAG, ZN |
 |
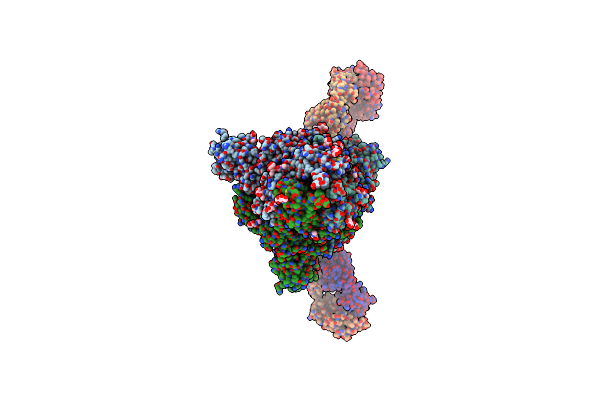
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
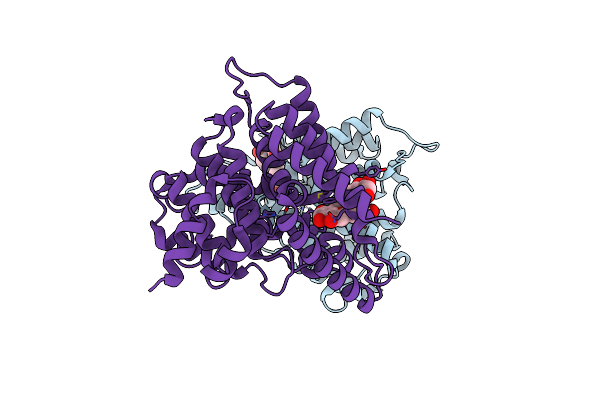
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: VIRAL PROTEIN/HYDROLASE/IMMUNE SYSTEM Ligands: NAG, ZN |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-06-12 Classification: HYDROLASE Ligands: ZN, MG, A1D6Q |
 |
Organism: Staphylococcus delphini
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-02-28 Classification: VIRAL PROTEIN |
 |
Organism: Staphylococcus delphini
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2024-02-28 Classification: VIRAL PROTEIN |
 |
Organism: Staphylococcus aureus, Staphylococcus delphini
Method: ELECTRON MICROSCOPY Release Date: 2024-02-28 Classification: VIRAL PROTEIN |
 |
Organism: Staphylococcus delphini
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2024-02-28 Classification: VIRAL PROTEIN |
 |
Organism: Staphylococcus aureus, Staphylococcus delphini
Method: ELECTRON MICROSCOPY Release Date: 2024-02-28 Classification: VIRAL PROTEIN |

