Search Count: 121
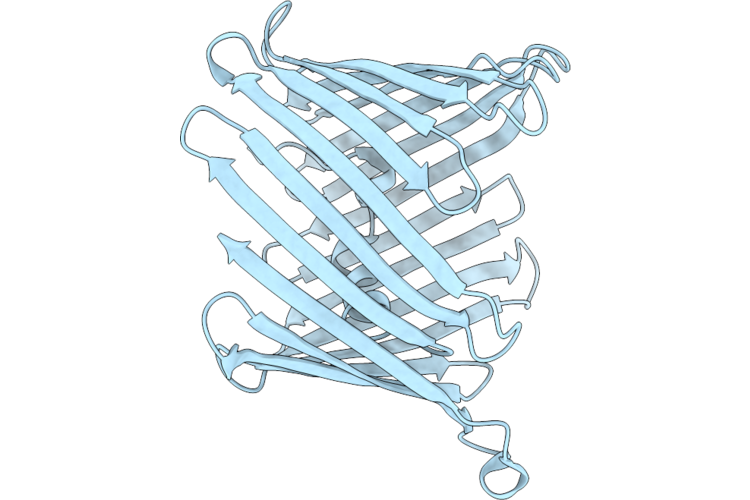 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
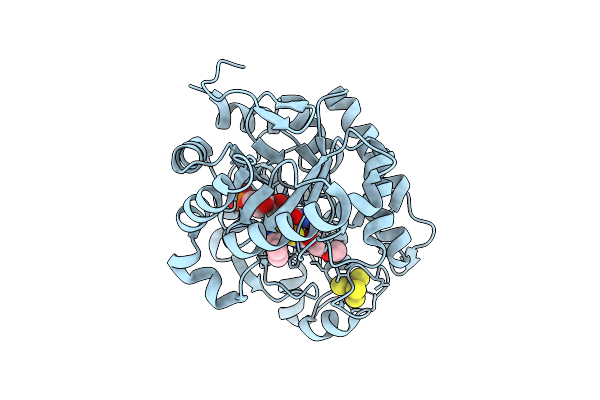 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: SF4, A1ELE, FMN |
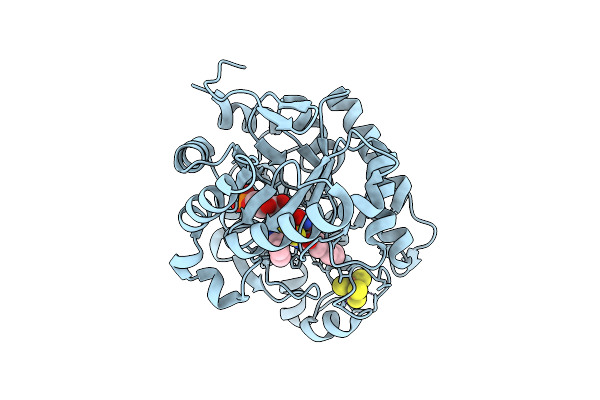 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 21
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELF |
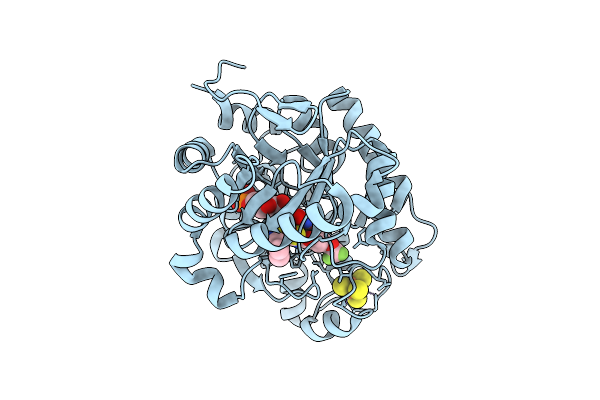 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 11
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELG |
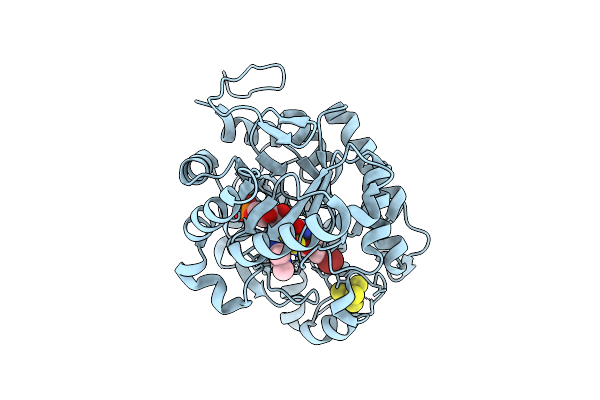 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 35
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELH |
 |
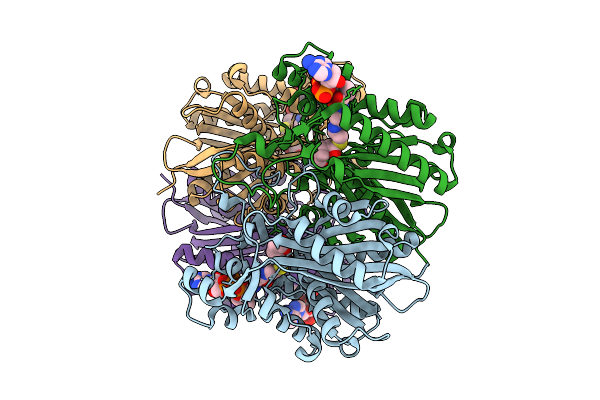
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 47
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1EMD |
 |
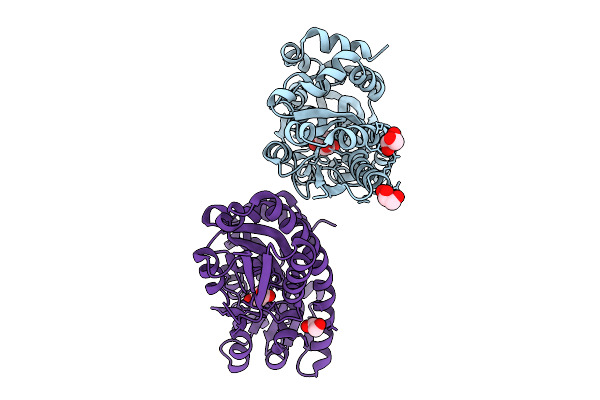
Crystal Structure Of Bioz C115S From Agrobacterium Tumefaciens In Complex With Galutaryl-Coa
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-10-29 Classification: BIOSYNTHETIC PROTEIN Ligands: GRA |
 |
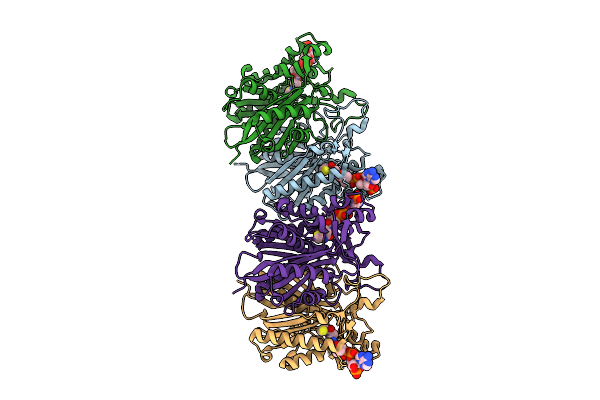
Crystal Structure Of Pa0884, The Sbp Component Of A Pseudomonas Aeruginosa Pao1 Tripartite Atp-Independent Periplasmic (Trap) Transporter, Complexed With Itaconate
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-09-10 Classification: TRANSPORT PROTEIN Ligands: ITN, GOL, EDO |
 |
Crystal Structure Of Bioz From Agrobacterium Tumefaciens In Complex With Galutaryl-Coa
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-08-20 Classification: BIOSYNTHETIC PROTEIN Ligands: COA |
 |
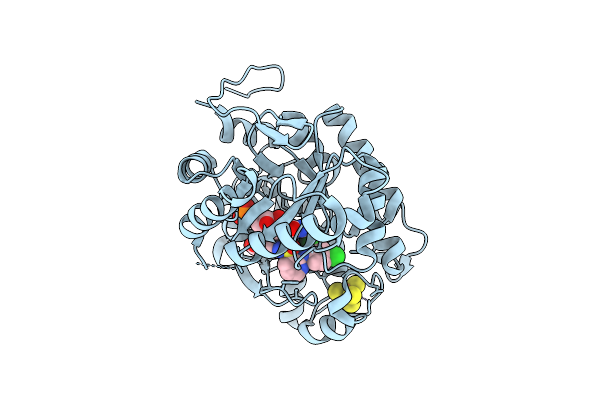
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1991
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-20 Classification: BIOSYNTHETIC PROTEIN Ligands: A1ECY, FMN, SF4, MG |
 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1872
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-08-20 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1EEQ |
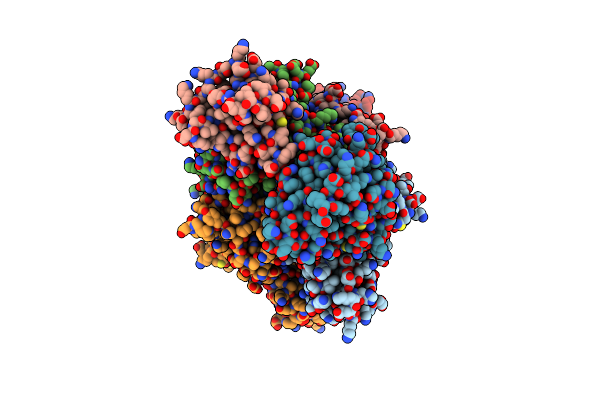 |
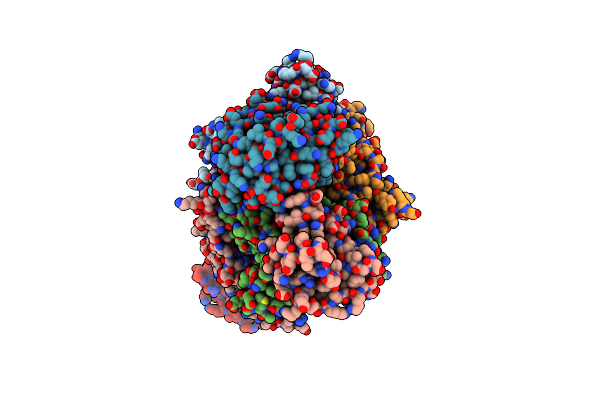
Cryo-Em Structure Of The Helicobacter Pylori Oordabc Complex In The Apo-Form
Organism: Helicobacter pylori
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: SF4, MG, TPP |
 |
Organism: Helicobacter pylori
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: SF4, A1D65, MG, TPP |
 |
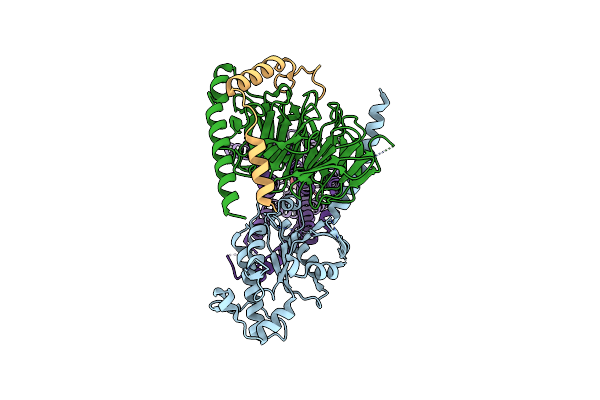
Crystal Structure Of Pa0884, The Sbp Component Of A Pseudomonas Aeruginosa Pao1 Tripartite Atp-Independent Periplasmic (Trap) Transporter, Complexed With Succinate
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-02-05 Classification: TRANSPORT PROTEIN Ligands: SIN |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#1 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#2 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#3 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#4 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#5 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |
 |
Cryoem Structure Of Beta-2-Adrenergic Receptor In Complex With Nucleotide-Free Gs Heterotrimer (#6 Of 20)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: G1I |

