Search Count: 30
All
Selected
 |
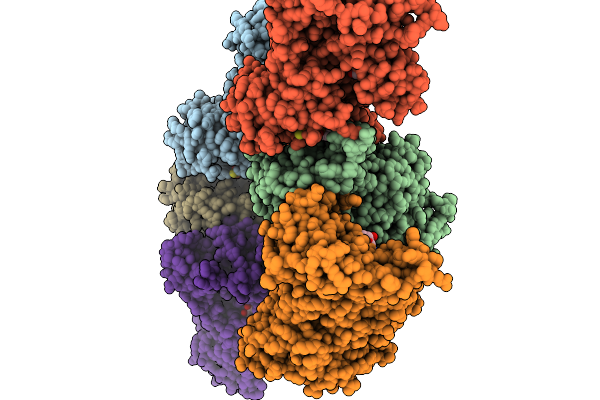
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP, GOL |
 |
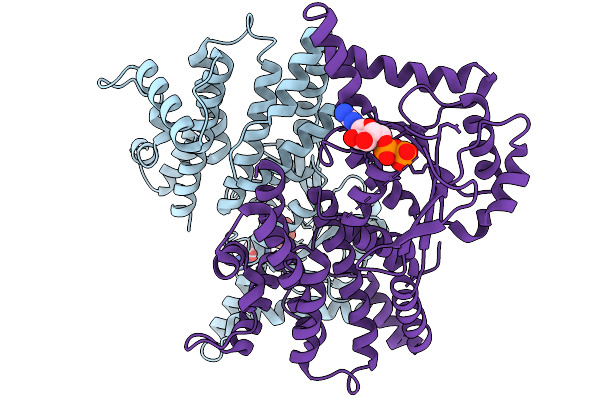
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GOL, GSP, MG |
 |
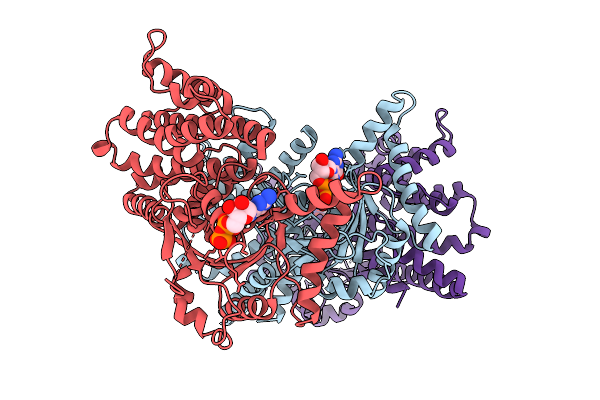
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.31 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP |
 |
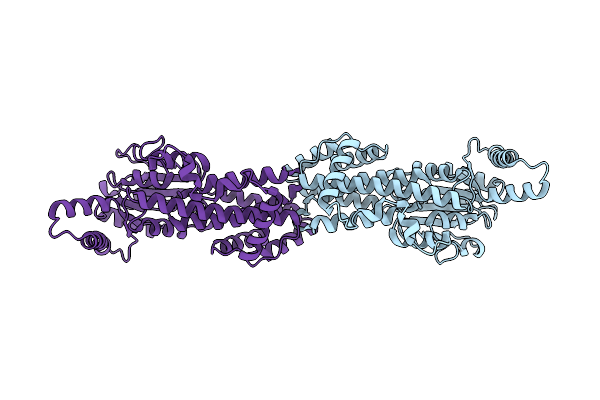
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN |
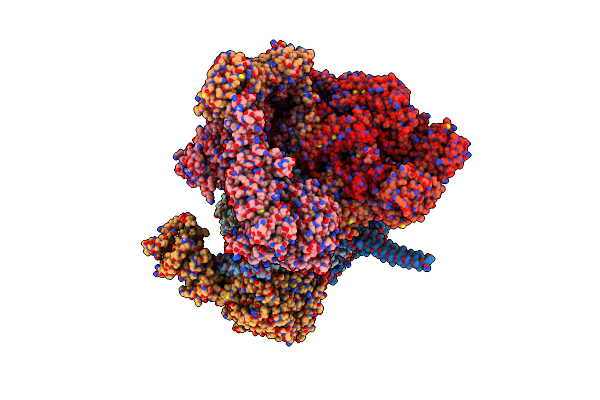 |
Structure Of Human Substrate-Free 26S Proteasome In The Presence Of Atpgs And Mg-132,Sa-Like State (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: ATP, MG, ADP, LDZ, ZN |
 |
Structure Of Substrate Engaged Midn-Bound Human 26S Proteasome, Eb-Midn (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: LDZ, ATP, MG, ADP, ZN |
 |
Structure Of Substrate Engaged Midn-Bound Human 26S Proteasome, Eb Midn_Ubl State (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrates-Engaged Midn-Bound Human 26S Proteasome,Ed-Midn State (Composite Map)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: IMMUNE SYSTEM Ligands: ATP, MG, ADP, ZN, LDZ |
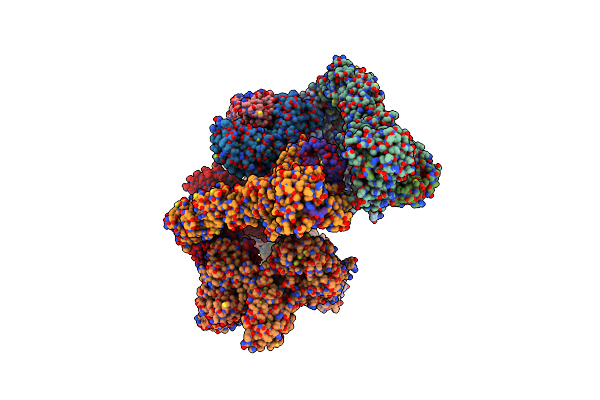 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
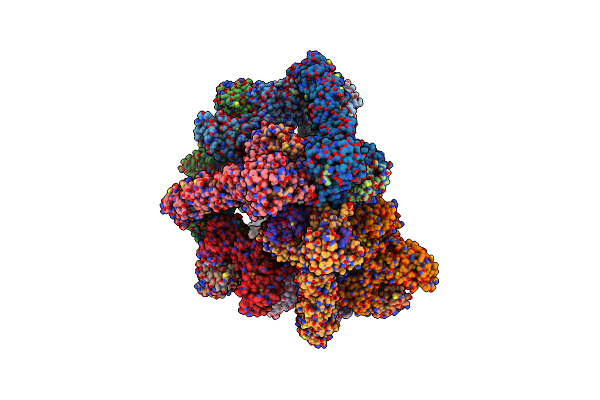 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
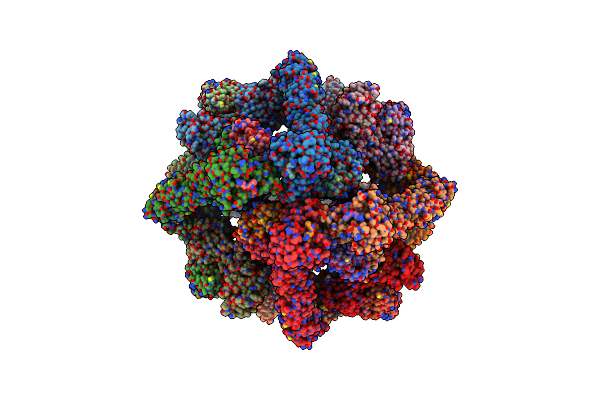 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-11 Classification: LIGASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-09-06 Classification: LIGASE Ligands: ZN |

