Search Count: 21
All
Selected
 |
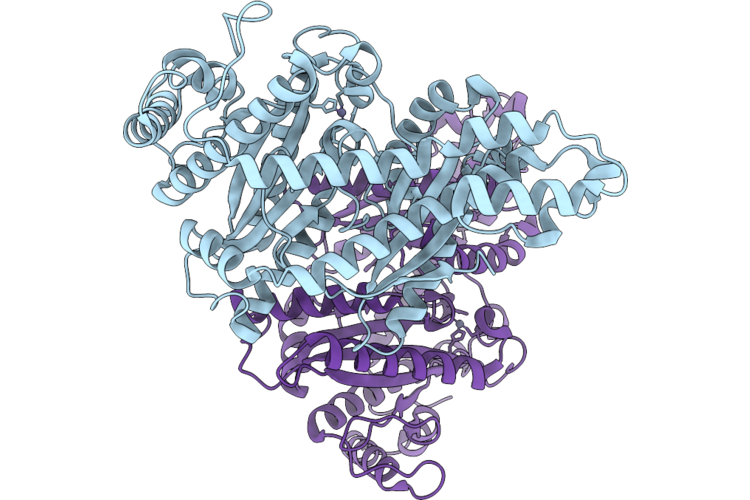
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
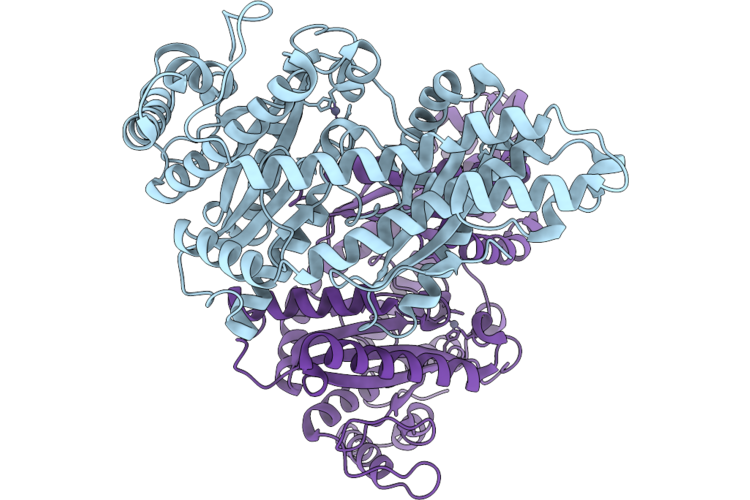
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 1
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 2
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.22 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
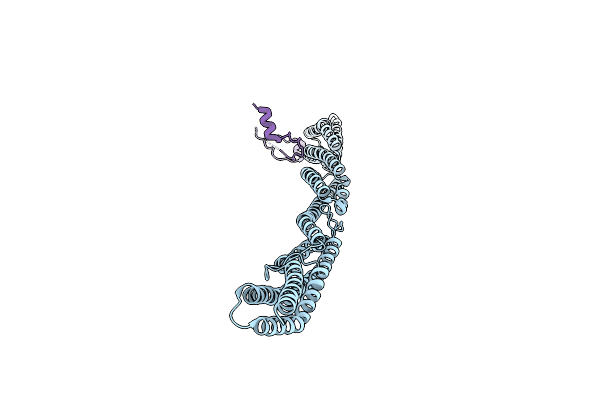 |
Crystal Structure Of Sarcomeric Protein Fatz-1 (Mini-Fatz-1 Construct) In Complex With Rod Domain Of Alpha-Actinin-2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2021-06-30 Classification: STRUCTURAL PROTEIN |
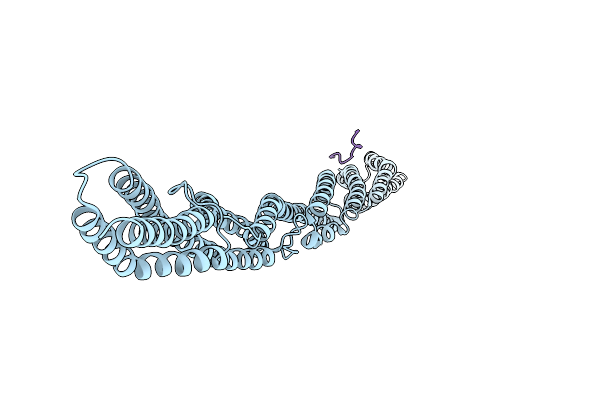 |
Crystal Structure Of Sarcomeric Protein Fatz-1 (D91-Fatz-1 Construct) In Complex With Rod Domain Of Alpha-Actinin-2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.80 Å Release Date: 2021-06-30 Classification: STRUCTURAL PROTEIN |
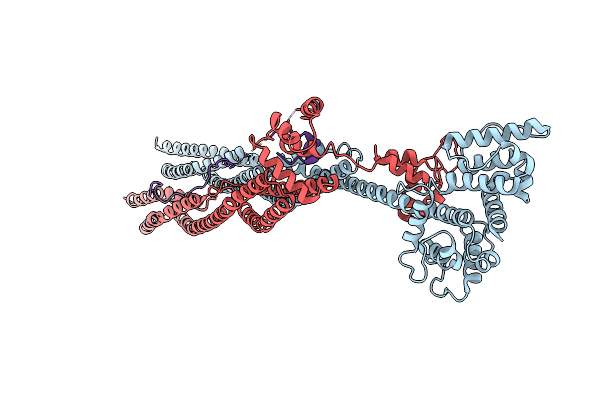 |
Crystal Structure Of Sarcomeric Protein Fatz-1 (D91-Fatz-1 Construct) In Complex With Half Dimer Of Alpha-Actinin-2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2021-06-30 Classification: STRUCTURAL PROTEIN |
 |
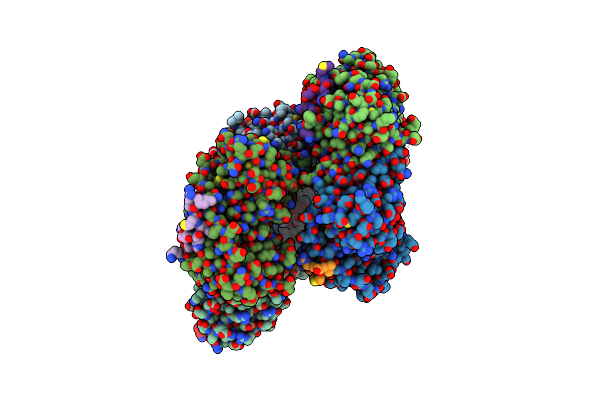
Papain Bound To A Natural Cysteine Protease Inhibitor From Streptomyces Mobaraensis
Organism: Carica papaya
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2019-12-04 Classification: HYDROLASE Ligands: EPE, N1W |
 |
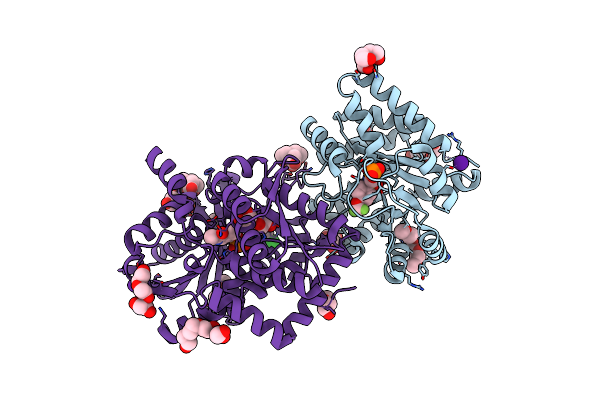
Crystal Structure Of Human Pcna In Complex With Parg (Poly(Adp-Ribose) Glycohydrolase) Peptide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2017-08-02 Classification: TRANSCRIPTION |
 |
The Crystal Structure Of Salmonella Typhimurium Tryptophan Synthase At 1.30A Complexed With N-(4'-Trifluoromethoxybenzenesulfonyl)-2-Amino-1-Ethylphosphate (F9) Inhibitor In The Alpha Site, Internal Aldimine
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2014-01-01 Classification: LYASE/LYASE INHIBITOR Ligands: F9F, BCN, CL, CS, PGE, PG5, EDO, PLP, PEG |
 |
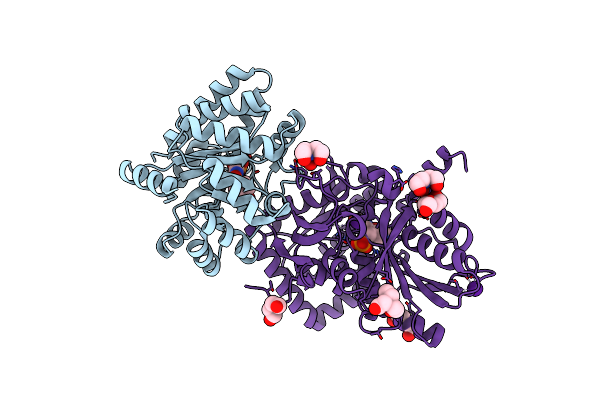
Crystal Structure Of Tryptophan Synthase From Salmonella Typhimurium With 2-Aminophenol Quinonoid In The Beta Site And The F6 Inhibitor In The Alpha Site
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2014-01-01 Classification: LYASE/LYASE INHIBITOR Ligands: F6F, NA, EDO, PEG, AQ3, PGE |
 |
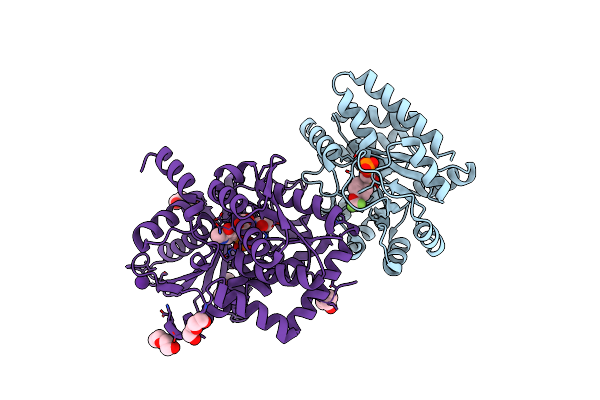
Tryptophan Synthase In Complex With Alpha Aminoacrylate E(A-A) Form And The F9 Inhibitor In The Alpha Site
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2013-12-25 Classification: LYASE/LYASE INHIBITOR Ligands: F9F, 0JO, PEG, BCN, CS |
 |
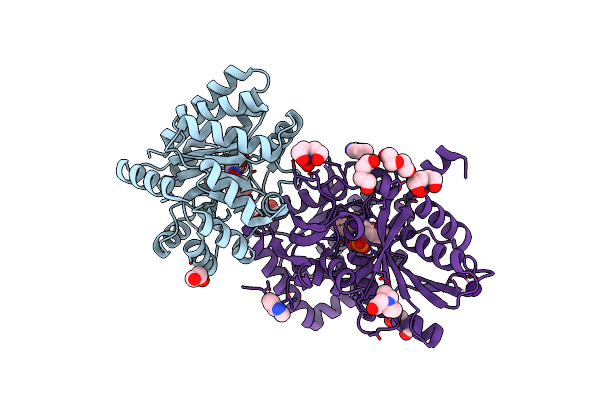
Crystal Structure Of Tryptophan Synthase At 1.45 A Resolution In Complex With 2-Aminophenol Quinonoid In The Beta Site And The F9 Inhibitor In The Alpha Site
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2013-12-25 Classification: LYASE/LYASE INHIBITOR Ligands: F9F, CL, 1D0, BCN, PEG, CS |
 |
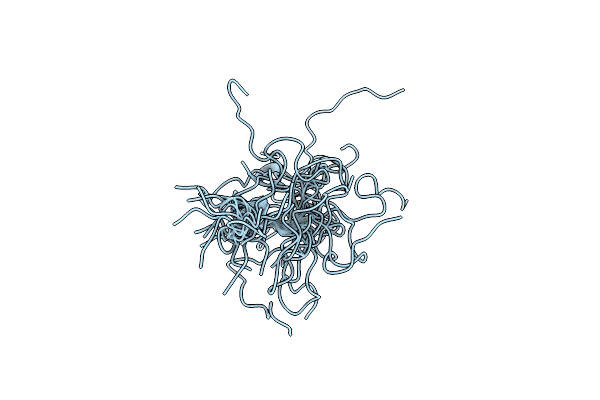
Crystal Structure Of Tryptophan Synthase At 1.65 A Resolution In Complex With Alpha Aminoacrylate E(A-A) And Benzimidazole In The Beta Site And The F9 Inhibitor In The Alpha Site
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2013-12-18 Classification: LYASE/LYASE INHIBITOR Ligands: F9F, PEG, BZI, 0JO, BCN, CS |
 |
Control Of K+ Channel Gating By Protein Phosphorylation: Structural Switches Of The Inactivation Gate, Nmr, 22 Structures
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 1999-04-27 Classification: POTASSIUM CHANNEL |

