Search Count: 514
All
Selected
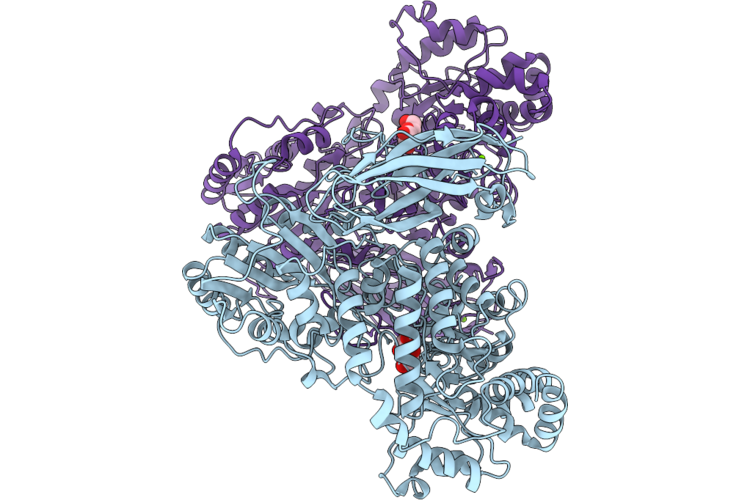 |
Crystal Structure Of Beta-Glucosidase From Bacteroides Ovatus Bound To Beta-D-Glucose
Organism: Bacteroides ovatus atcc 8483
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-05-20 Classification: HYDROLASE Ligands: BGC, MG |
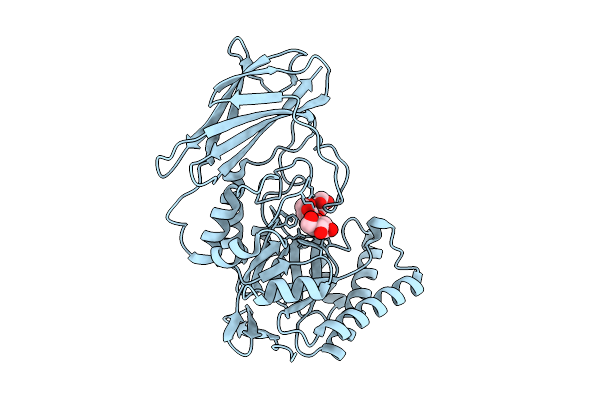 |
Crystal Structure Of Gh158(Pro) Soaked With Laminaritriose At 1.30 Angstrom Resolution.
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: BGC, CL |
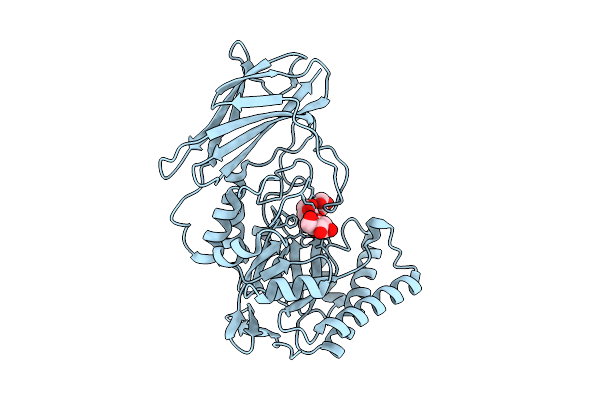 |
Crystal Structure Of Gh158(Pro) Soaked With Mixed Linkage (G3G4G) Oligosaccharide At 1.70 Angstrom Resolution
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: BGC, CL |
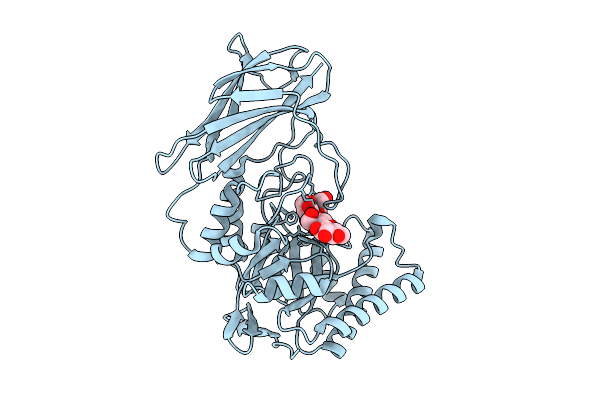 |
Crystal Structure Of Gh158(Pro) Soaked With Mixed Linkage (G4G3G4G) Oligosaccharide At 1.30 Angstrom Resolution
Organism: Metagenome
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: BGC, CL |
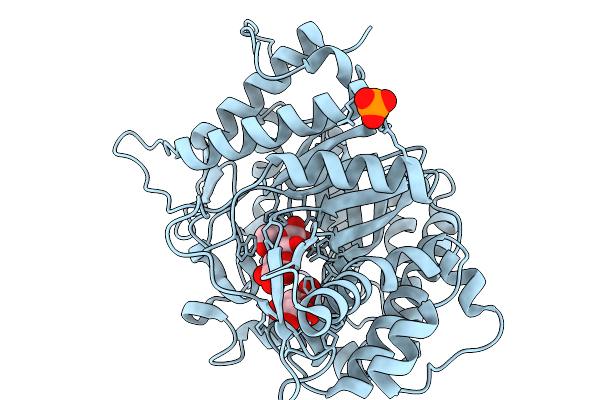 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, BGC, PO4 |
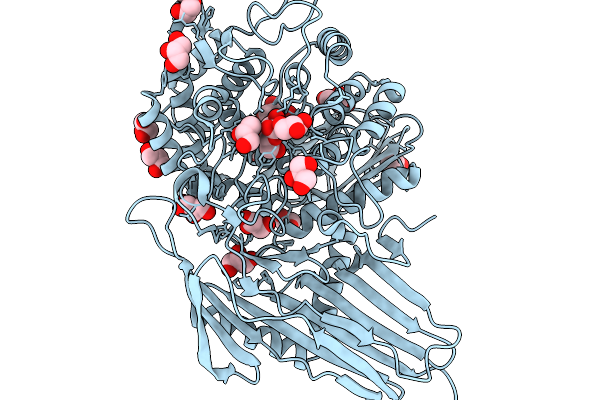 |
Crystal Structure Of The C-Terminally Truncated Human E574Q-Pgghg Mutant In Complex With Glucose
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2026-02-11 Classification: SUGAR BINDING PROTEIN Ligands: CL, GOL, BGC, GLC, PEG |
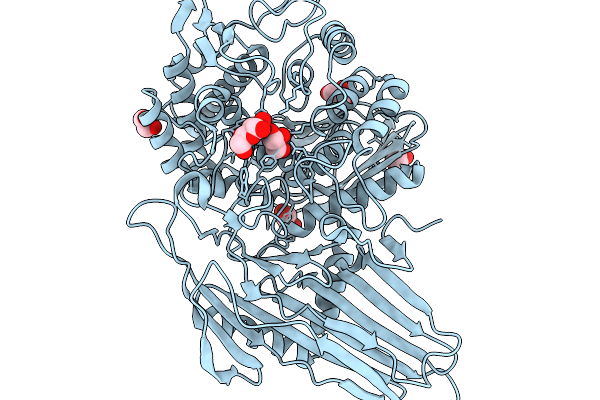 |
Crystal Structure Of C-Terminally Truncated Human Pgghg In Complex With Glucose
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-02-11 Classification: SUGAR BINDING PROTEIN Ligands: CL, GOL, BGC |
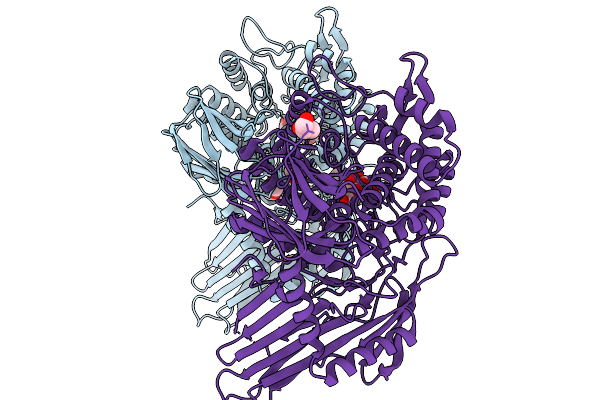 |
Crystal Structure Of C-Terminally Truncated Human Pgghg In Complex With Glucose.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-02-11 Classification: SUGAR BINDING PROTEIN Ligands: CL, GLC, BGC, GOL, PEG |
 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
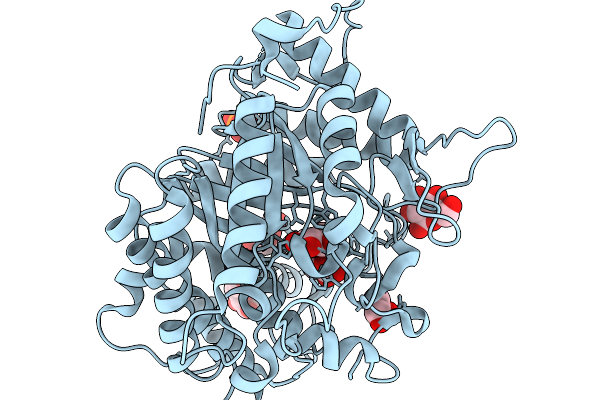
Crystal Structure Of Glucsoe Bound Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
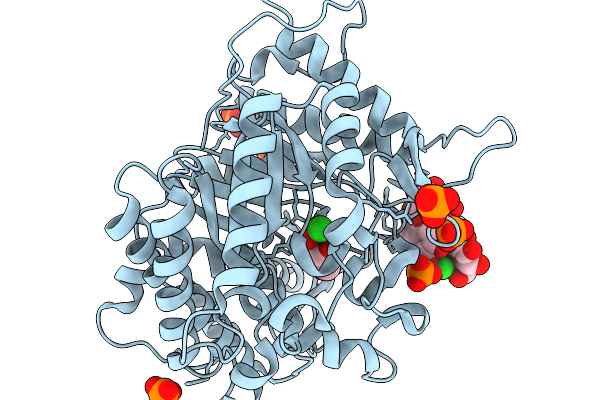
Crystal Structure Of Glucose Bound Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, GOL, PO4 |
 |
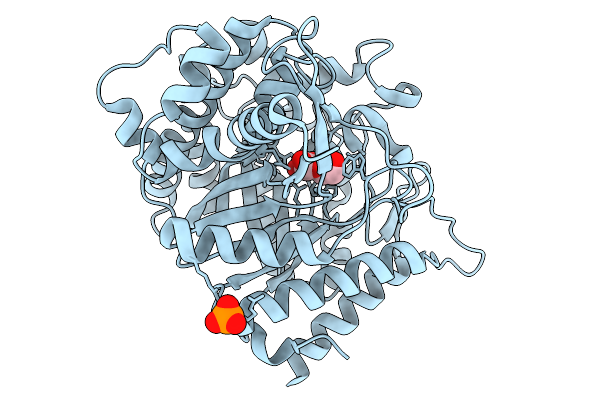
Crystal Structure Of Cellobiose And Glucose Bound Glucose Toleranant Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4, CL |
 |
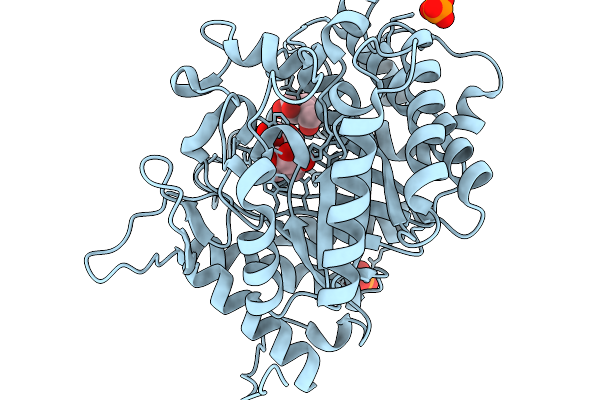
Crystal Structure Of Glucose Bound Highly Glucose Tolerant Gh1 Beta-Glcosidase Mutant (Unbgl1_C188V_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
Crystal Structure Of Glucose Bound Covalent Intermediate Of Gh1 Beta-Glucosidase (Unbgl1)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
Organism: Thermoproteus sp. az2
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-14 Classification: HYDROLASE Ligands: BGC, GOL, ACT |
 |
Organism: Gut metagenome
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: HYDROLASE Ligands: PO4, BGC |
 |
Spitrobot-2 Advances Time-Resolvedcryo-Trapping Crystallography To Under 25 Ms: Xylose Isomerase Bound With Glucose (25 Ms Soaking)
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-03 Classification: ISOMERASE Ligands: MG, MN, BGC |
 |
Spitrobot-2 Advances Time-Resolvedcryo-Trapping Crystallography To Under 25 Ms: Xylose Isomerase Bound With Glucose (50 Ms Soaking)
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-03 Classification: ISOMERASE Ligands: MG, MN, BGC |
 |
Crystal Structure Of Tegh116 From Thermosynechococcus Elongatus With Glucose
Organism: Thermosynechococcus vestitus bp-1
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-11-26 Classification: HYDROLASE Ligands: BGC, GOL, CA, TLA |
 |
F420-Dependent Glucose-6-Phosphate Dehydrogenase From Thermomicrobium Roseus With Glucose
Organism: Thermomicrobium roseum dsm 5159
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2025-11-19 Classification: OXIDOREDUCTASE Ligands: BGC, NA |

