Search Count: 6
All
Selected
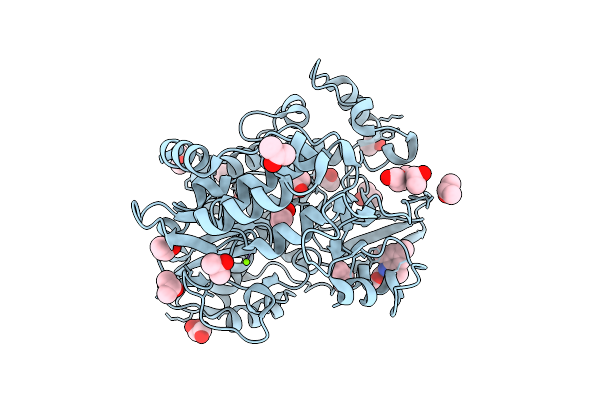 |
Pandda Analysis Group Deposition -- Crystal Structure Of Enterovirus D68 3Dpol In Complex With Z57260516
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: IPA, PEG, GOL, B0V, MG |
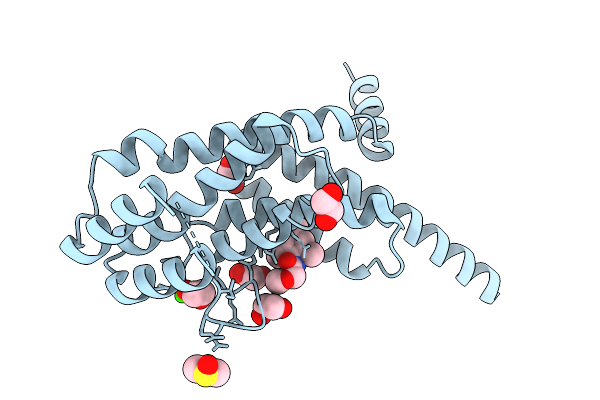 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-11-19 Classification: PROTEIN BINDING Ligands: B0V, EDO, CA, DMS |
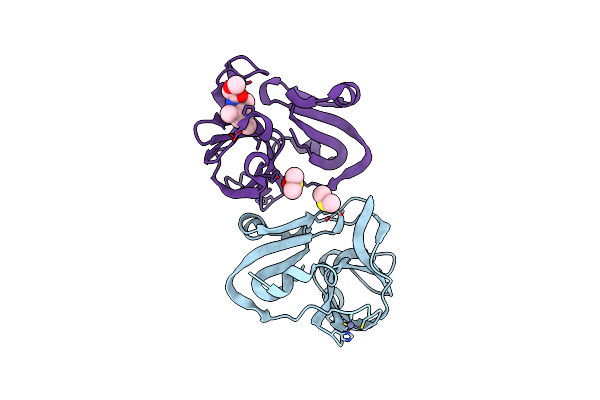 |
Group Deposition For Crystallographic Fragment Screening Of Coxsackievirus A16 (G-10) 2A Protease -- Crystal Structure Of Coxsackievirus A16 (G-10) 2A Protease In Complex With Z57260516 (A71Ev2A-X0387)
Organism: Coxsackievirus a16
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2024-04-24 Classification: HYDROLASE Ligands: ZN, DMS, B0V |
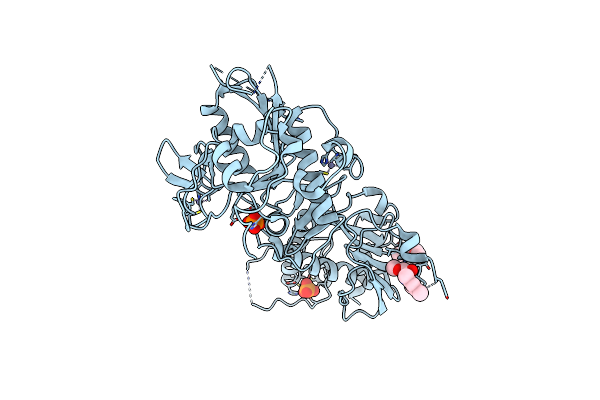 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp14 In Complex With Z57260516
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2022-03-16 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: ZN, PO4, B0V |
 |
Pandda Analysis Group Deposition Of Models With Modelled Events (E.G. Bound Ligands) -- Crystal Structure Of Nudt7 In Complex With Fmopl000710A
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2019-03-27 Classification: HYDROLASE Ligands: ACT, DMS, B0V |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Dclre1A In Complex With Fmopl000710A
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2018-08-08 Classification: CELL CYCLE Ligands: MLI, NI, B0V |

