Search Count: 16
All
Selected
 |
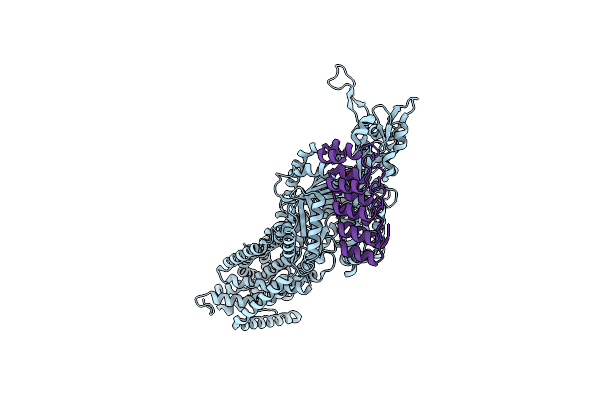
Crystal Structure Of Multidrug Efflux Transporter Oqxb From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN Ligands: LMT, GOL |
 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN Ligands: MIY |
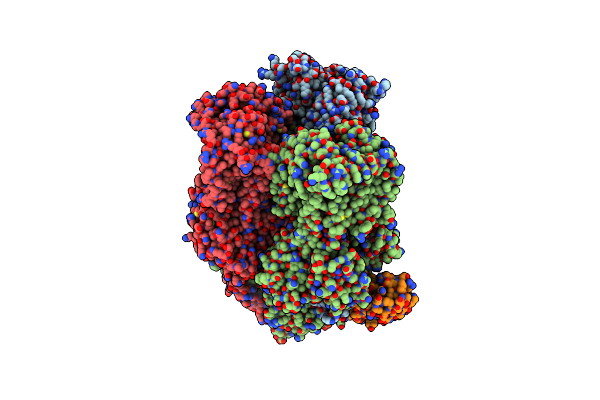 |
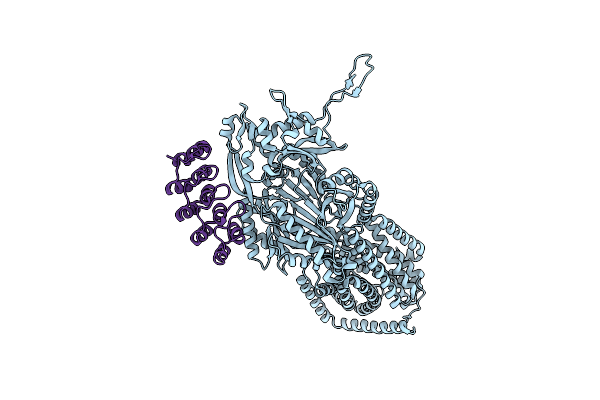
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
 |
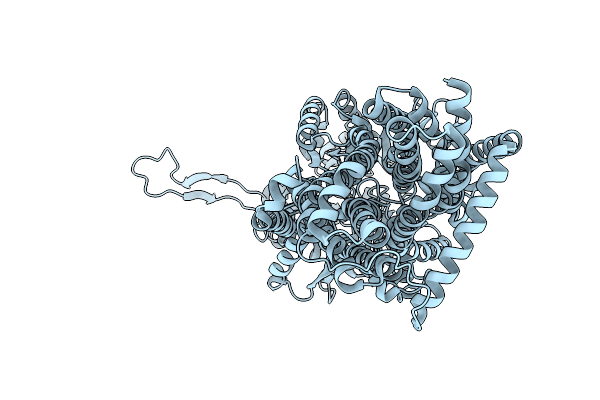
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.55 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Single Particle Cryo-Em Structure Of The Multidrug Efflux Pump Oqxb From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Synthetic construct, Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN Ligands: MIY |
 |
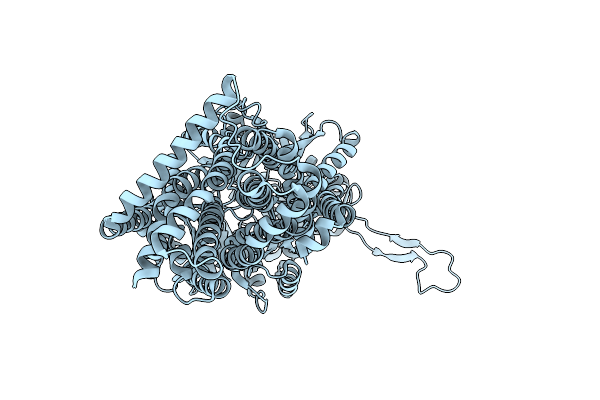
Organism: Synthetic construct, Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
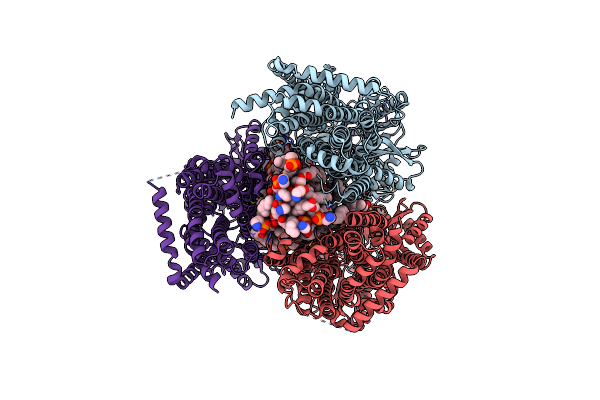 |
Single Particle Cryo-Em Structure Of The Multidrug Efflux Pump Mdtf From Escherichia Coli
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: PTY, D12 |
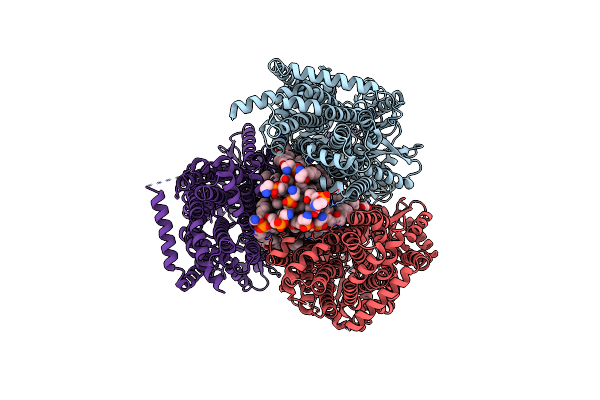 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: PTY, D12 |
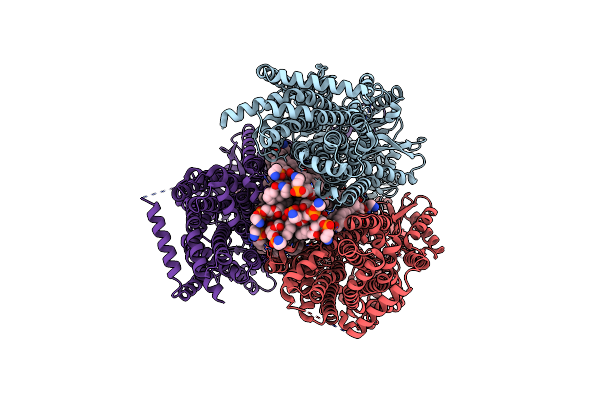 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: D12, PTY, RHQ |
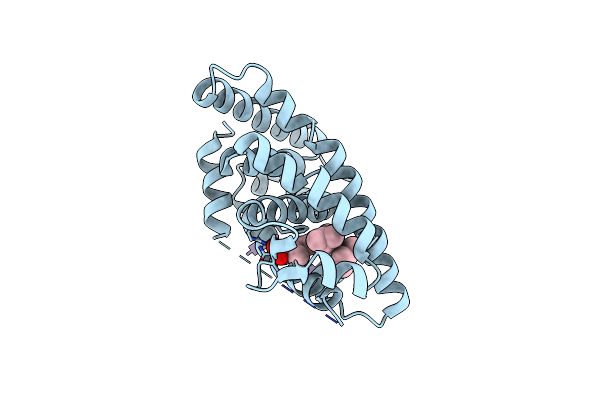 |
Crystal Structure Of Human Retinoid X Receptor Alpha Ligand Binding Domain Complex With Uab116 And Coactivator Peptide Grip-1
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2023-03-15 Classification: TRANSCRIPTION Ligands: OI2 |
 |
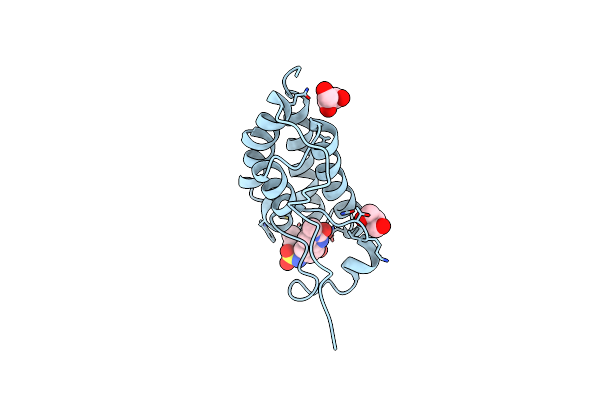
Crystal Structure Of Human Retinoid X Receptor Alpha Ligand Binding Domain Complex With Uab113 And Coactivator Peptide Grip-1
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-03-15 Classification: TRANSCRIPTION Ligands: OI5 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2022-08-31 Classification: GENE REGULATION Ligands: EM0, GOL |

