Search Count: 5,125
All
Selected
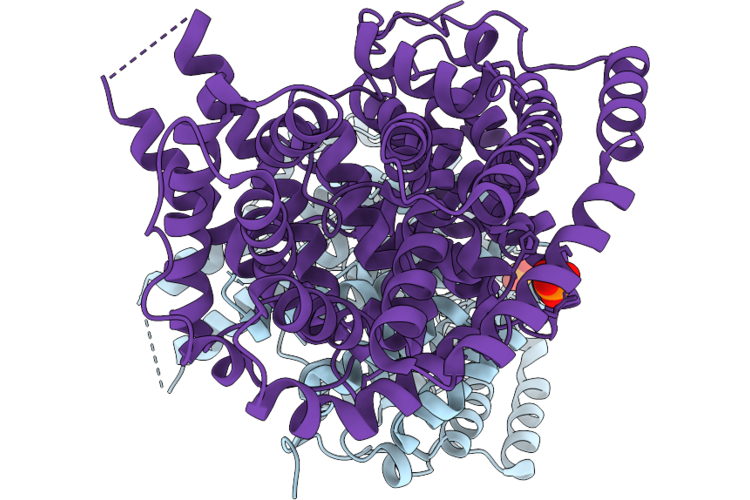 |
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-05-13 Classification: LYASE Ligands: PO4 |
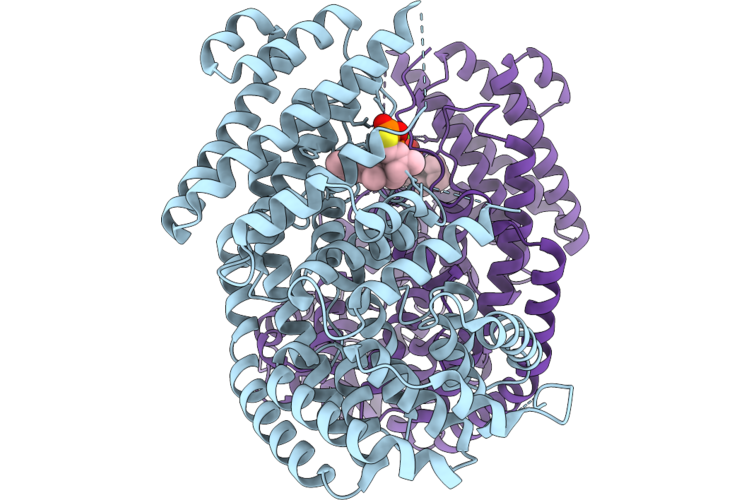 |
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2026-05-13 Classification: LYASE Ligands: MG, FPS |
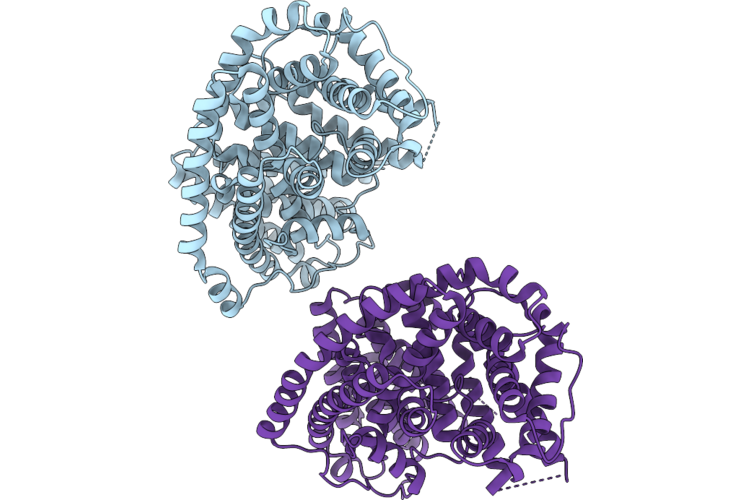 |
Crystal Structure Of Bicyclogermacrene Synthase Mutant I290V/I385C/V434C/L454C/V476W/L558I
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-05-13 Classification: LYASE |
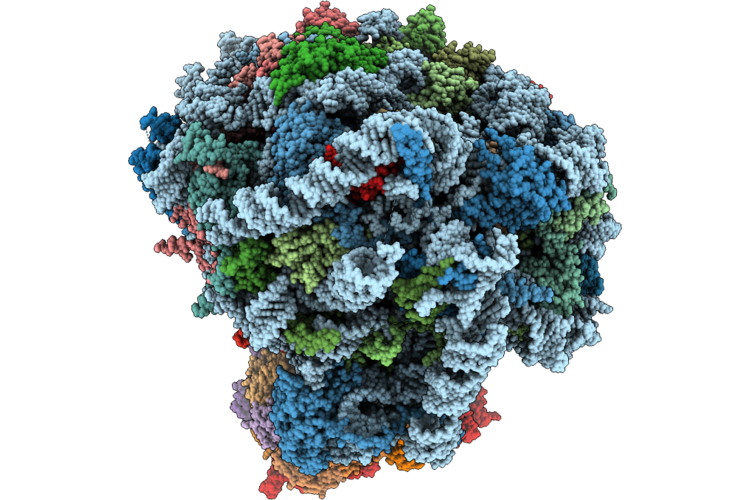 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Staphylococcus aureus subsp. aureus nctc 8325
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: GTP, GOL |
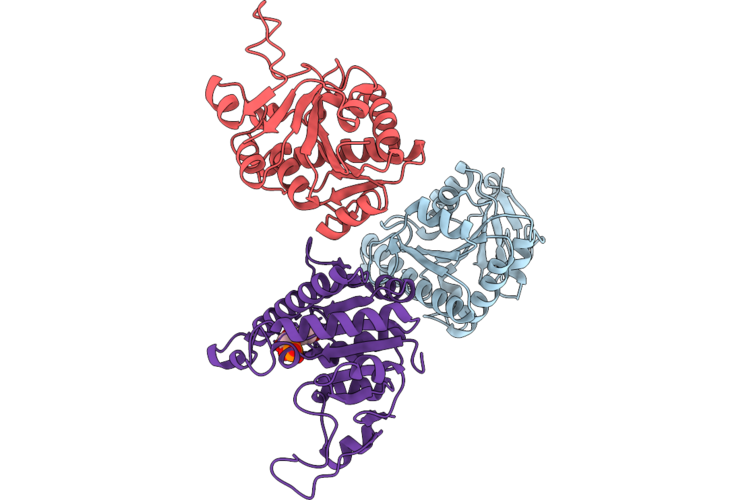 |
Organism: Staphylococcus aureus subsp. aureus nctc 8325
Method: X-RAY DIFFRACTION Resolution:3.26 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: U5P |
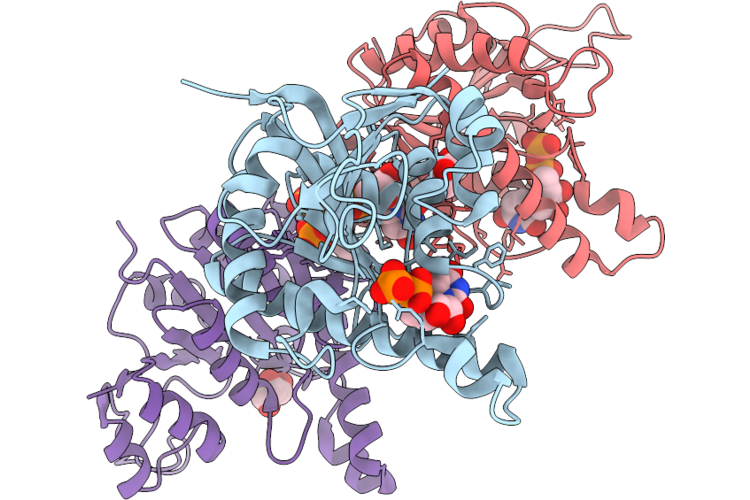 |
Organism: Staphylococcus aureus subsp. aureus nctc 8325
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: UDP, GOL, PO4 |
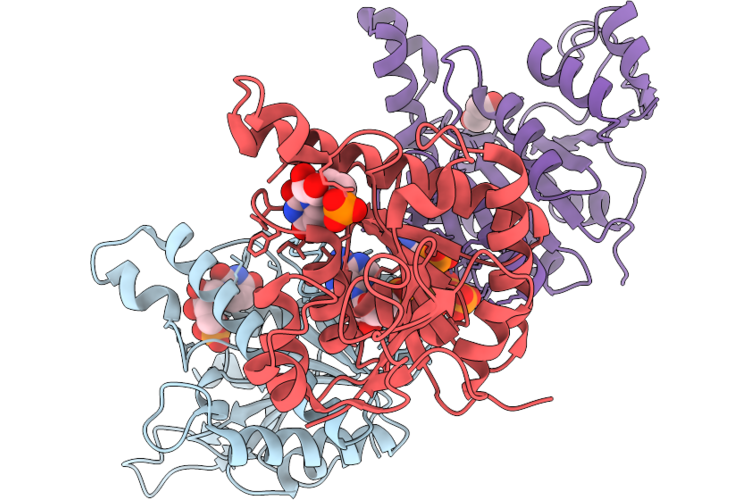 |
Organism: Staphylococcus aureus subsp. aureus nctc 8325
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: U5P, ATP, PO4, GOL |
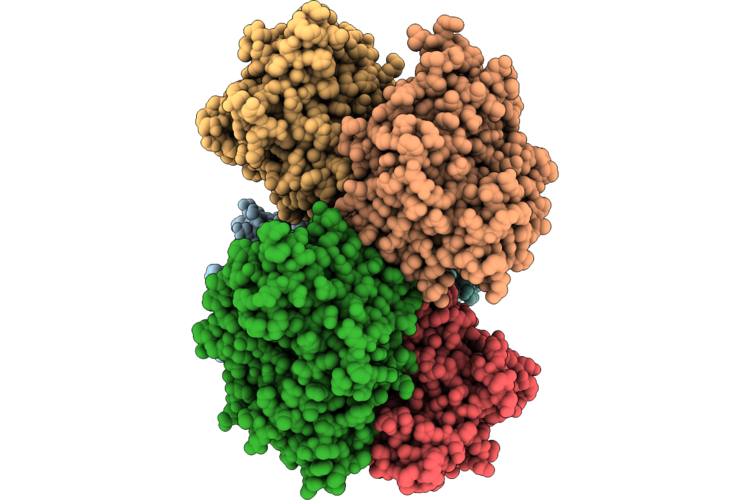 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.74 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
 |
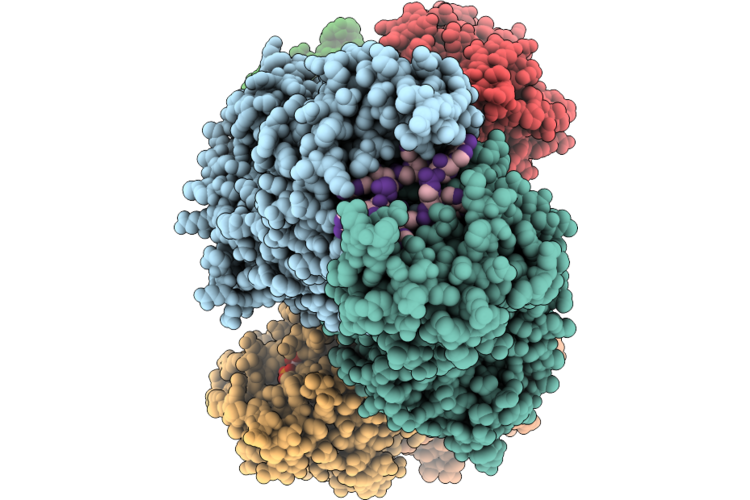
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
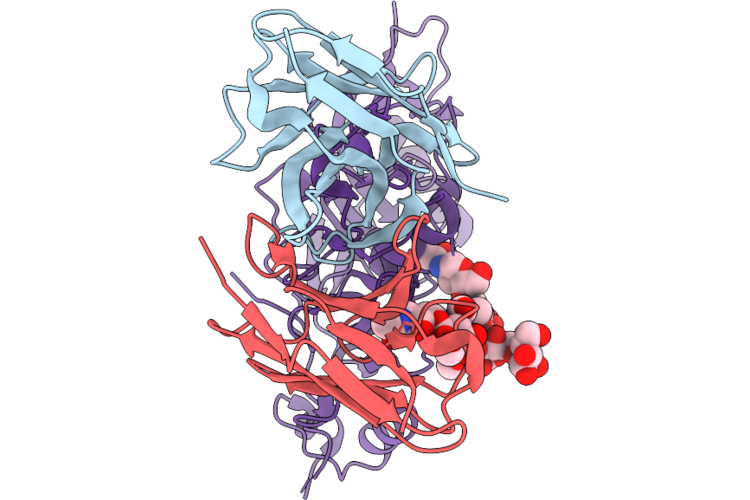 |
Epitope And Functional Classification Of Human Neutralizing Antibodies Against Sftsv Gn
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN |
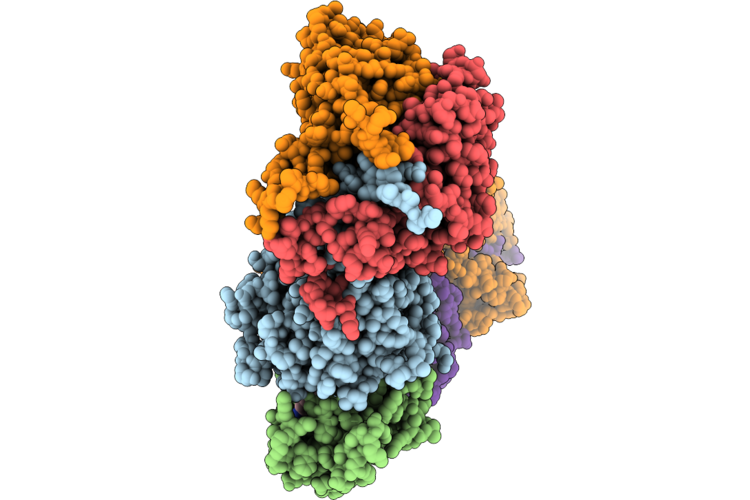 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10088
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
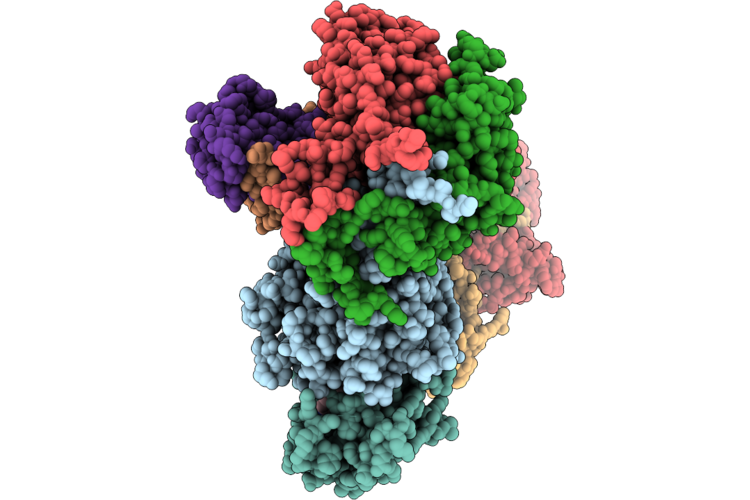 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2, Unc10088 And Bach1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10415667
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J21 |

