Search Count: 127
All
Selected
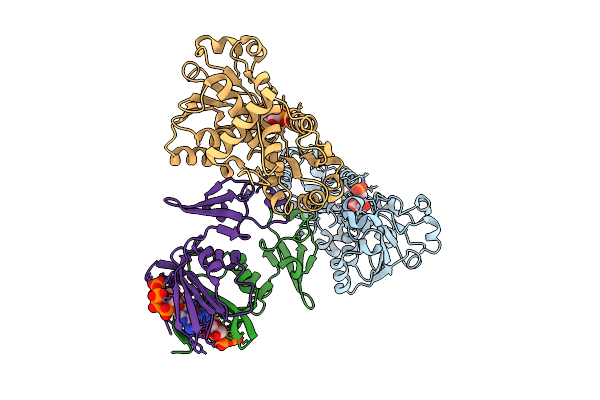 |
Crystal Structure Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Pala, Atp, Gtp, And Mg2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: PAL, ZN, ATP, MG, GTP |
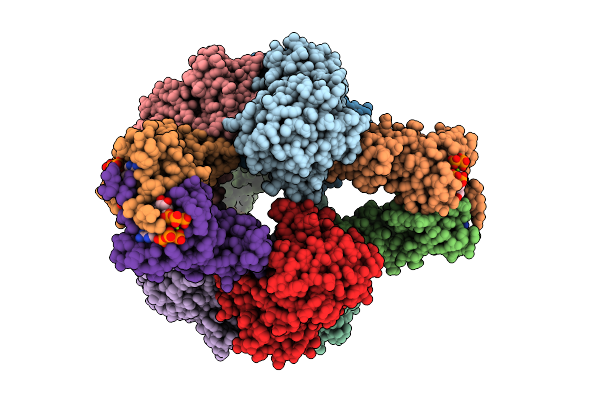 |
Crystal Structure Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, And Mg2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, MG |
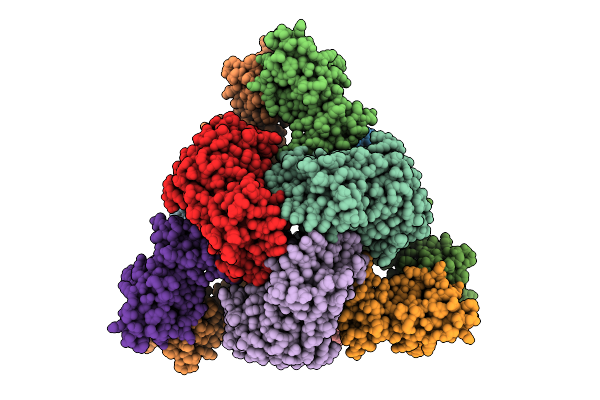 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The Ligand-Free T-State
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN |
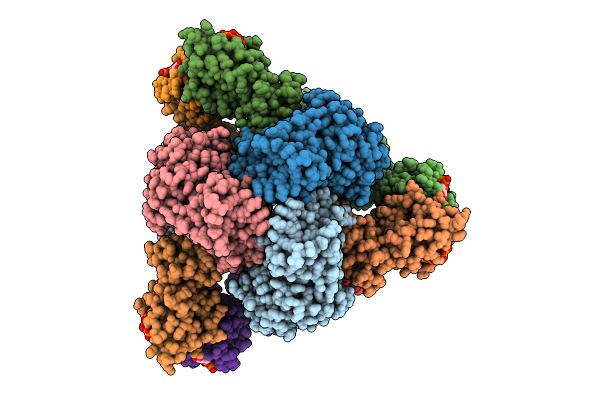 |
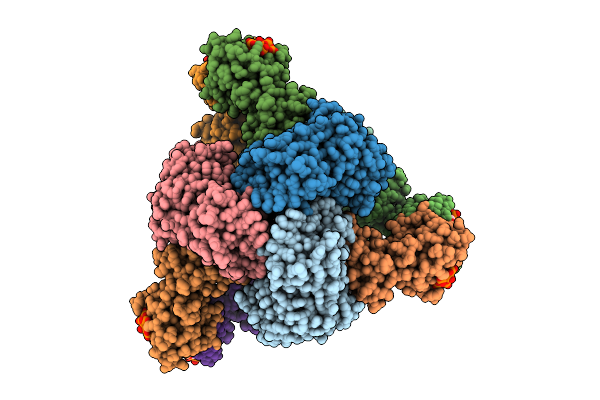
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Ctp, Utp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, CTP, UTP, MG, CP |
 |
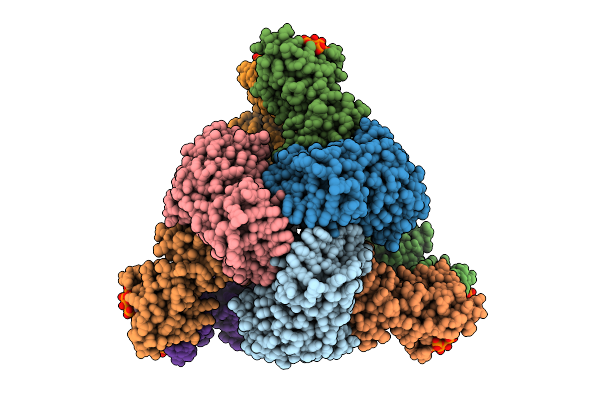
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Ctp, Utp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, CTP, UTP, MG |
 |
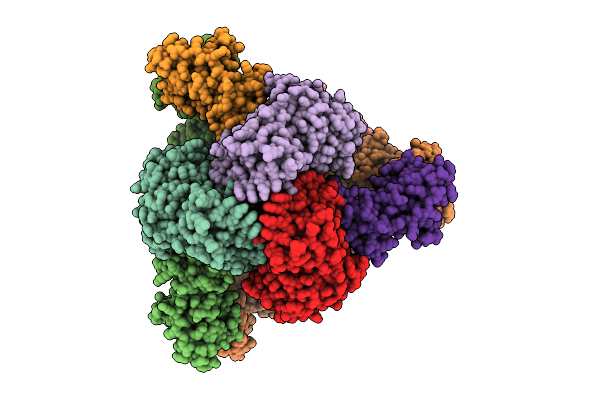
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Ctp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, CTP, MG |
 |
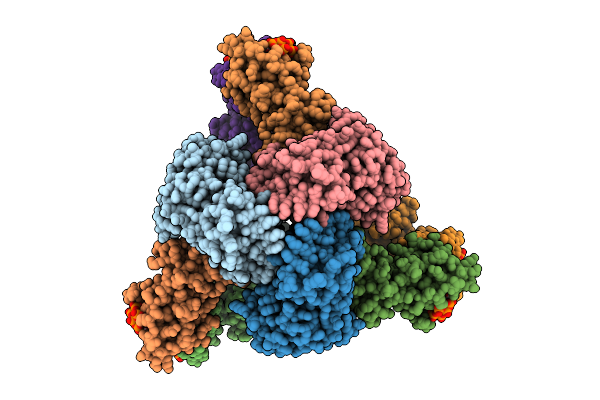
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp And Succinate
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN |
 |
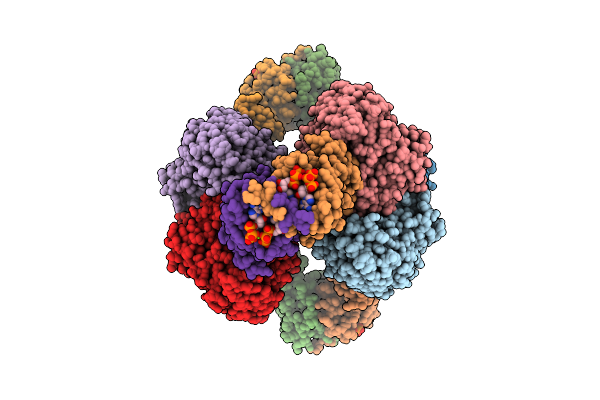
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, MG |
 |
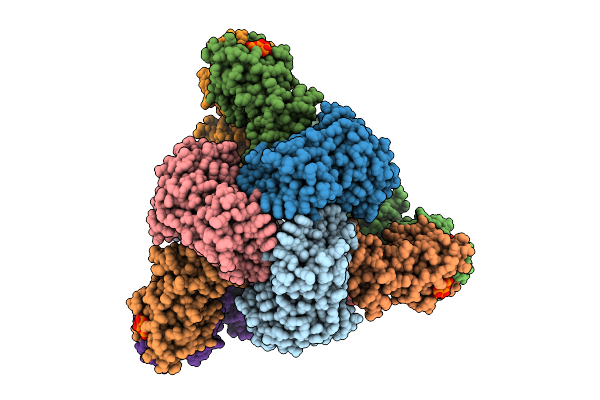
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, Gtp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, GTP, MG |
 |
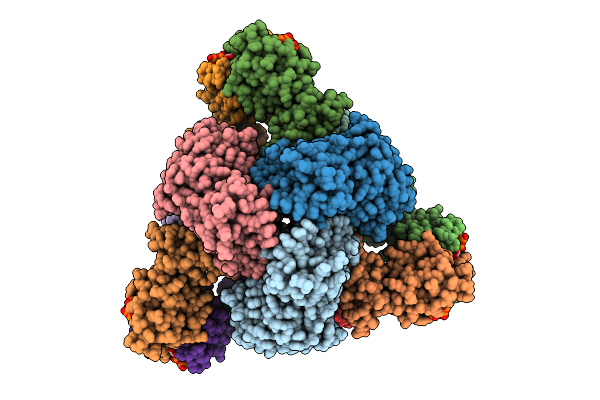
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In An Expanded State Complexed With Cp, Atp, Gtp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, ATP, GTP, MG, CP |
 |
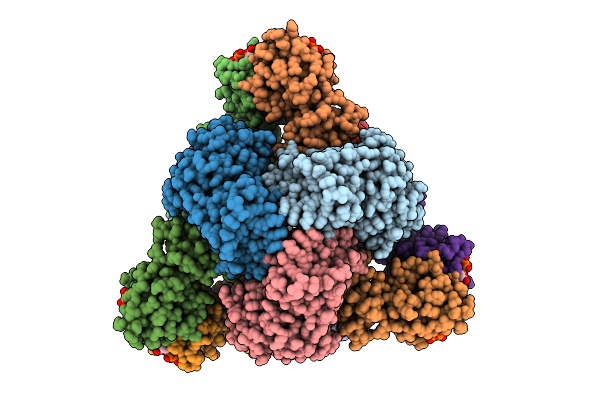
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Atp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, ATP, MG, CP |
 |
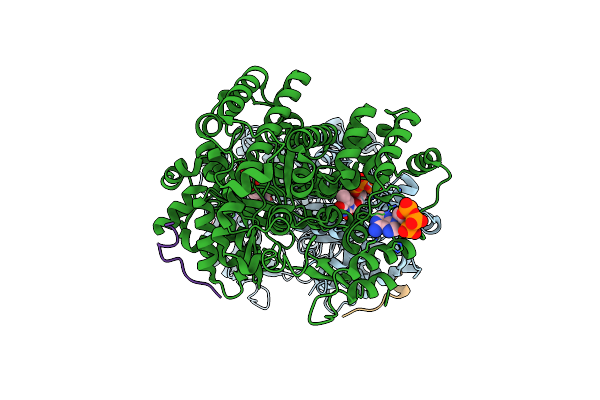
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Ctp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: CYTOSOLIC PROTEIN Ligands: ZN, CTP, MG, CP |
 |
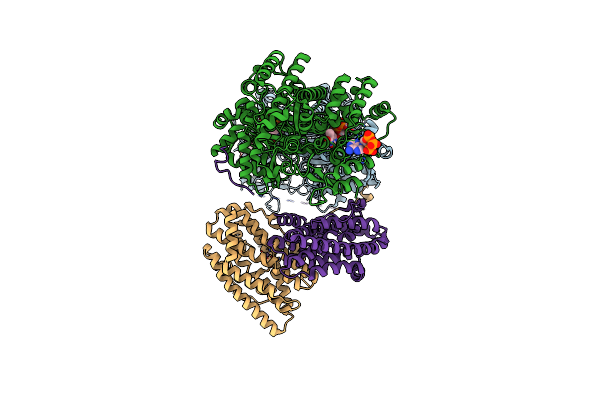
Consensus Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP |
 |
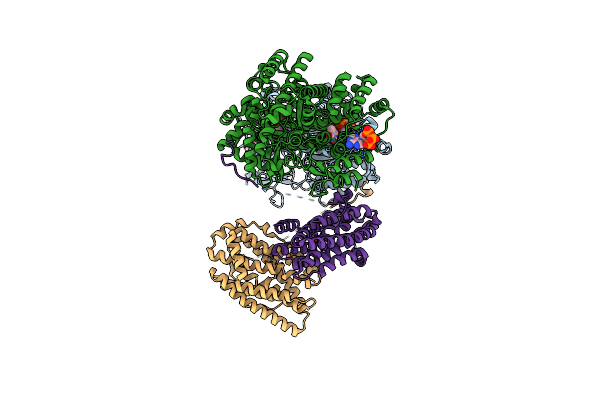
Consensus Full-Complex Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
 |
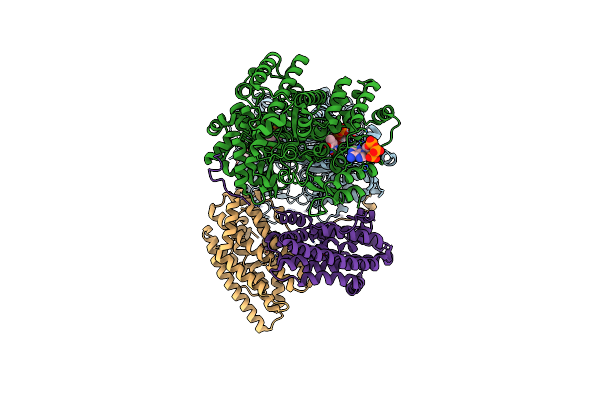
Class 2 Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
 |
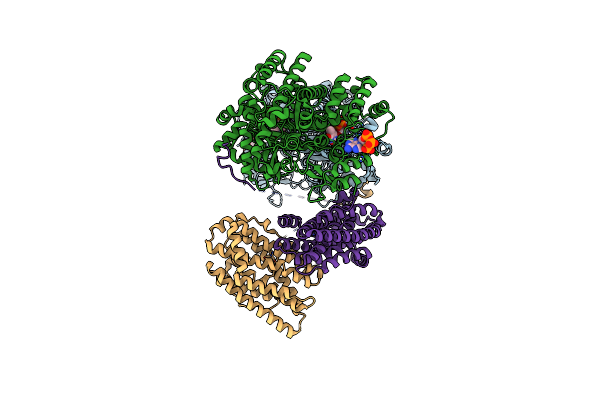
Class 5 Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
 |
Class 9 Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
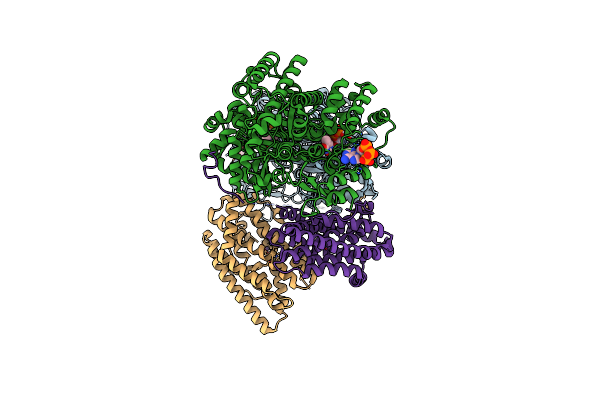 |
Class 12 Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
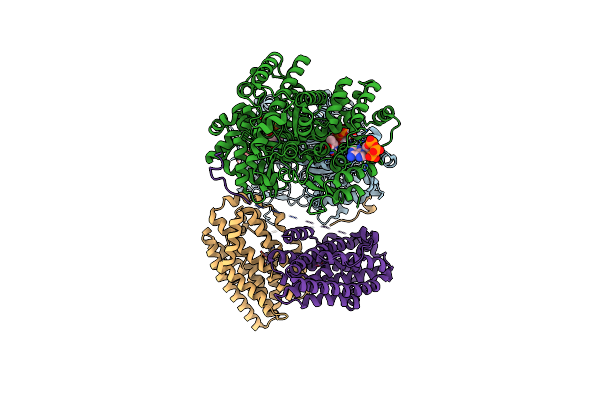 |
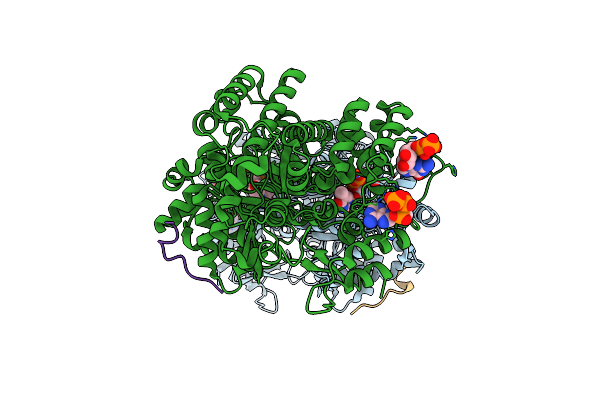
Class 15 Model For Preturnover Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, MN |
 |
Consensus Model For Pre-Reduction Condition Of Bacillus Subtilis Ribonucleotide Reductase Complex
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: OXIDOREDUCTASE Ligands: TTP, MG, ATP, GDP |

