Search Count: 83
All
Selected
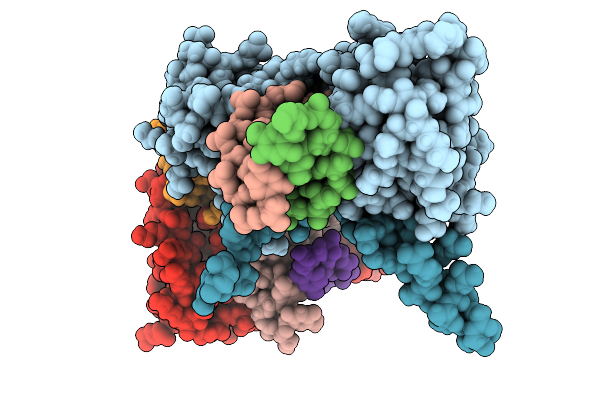 |
Structure Of Pou2F3 Pou Domains Bound To Coactivator Oca-T2 And Dna (2.8 Angstrom Resolution)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2025-10-01 Classification: TRANSCRIPTION |
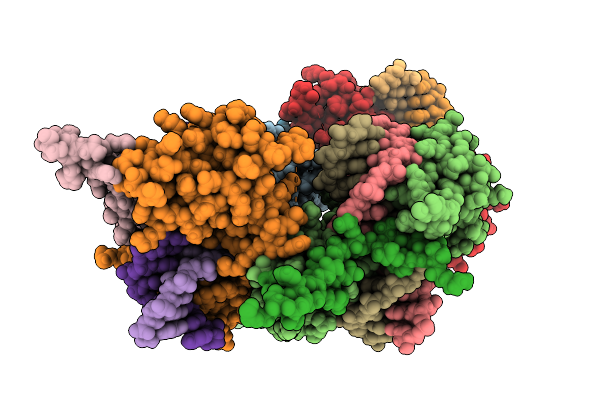 |
Structure Of Pou2F3 Pou Domains Bound To Coactivator Oca-T2 And Dna (2.1 Angstrom Resolution)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-10-01 Classification: TRANSCRIPTION |
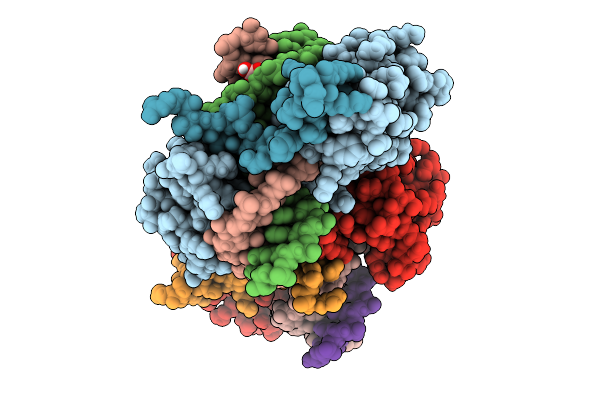 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-10-01 Classification: TRANSCRIPTION Ligands: BO3 |
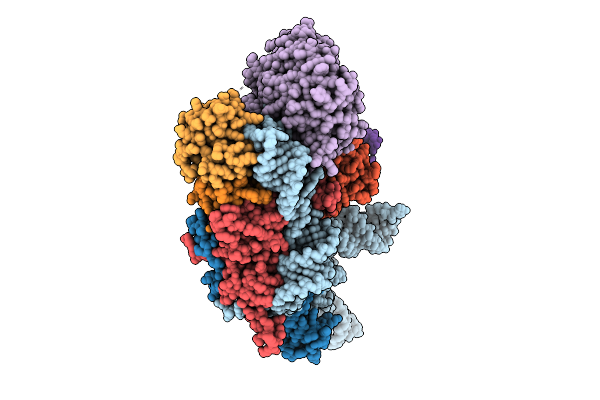 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-07-16 Classification: RNA BINDING PROTEIN |
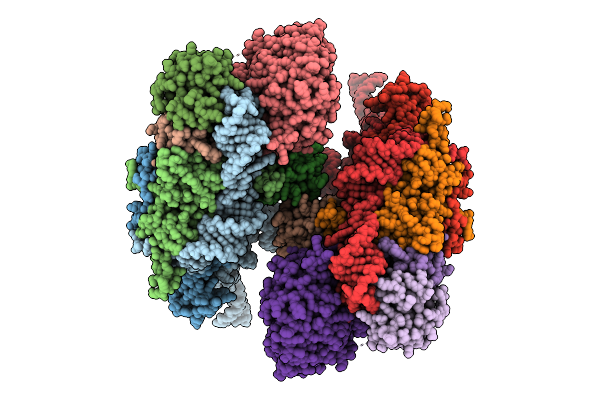 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2025-07-16 Classification: RNA BINDING PROTEIN |
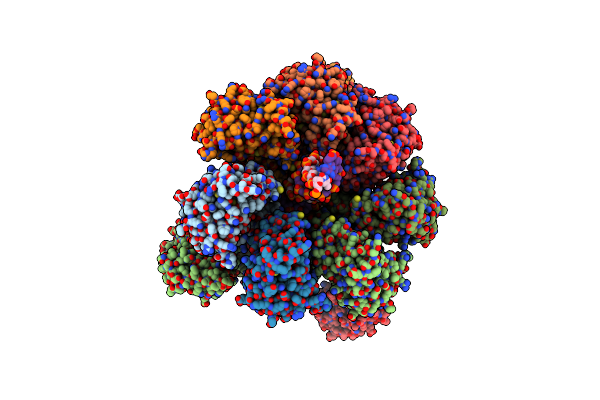 |
Organism: Adeno-associated virus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-02-12 Classification: VIRAL PROTEIN/DNA |
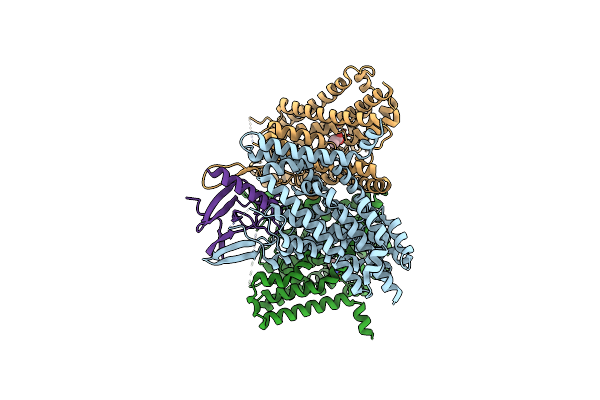 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-05-01 Classification: PROTEIN TRANSPORT Ligands: ALA |
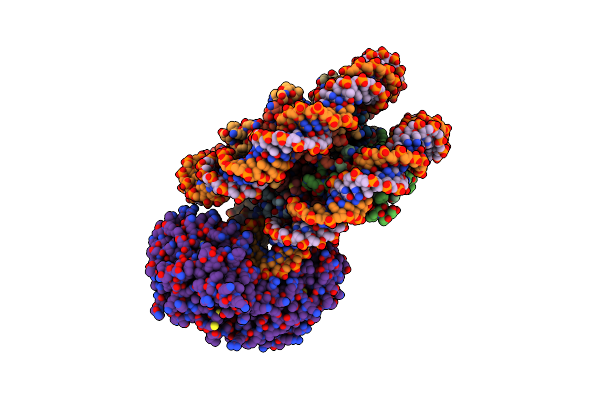 |
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: GENE REGULATION |
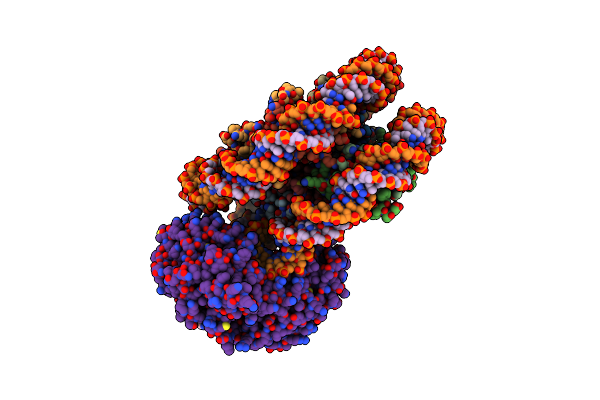 |
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: GENE REGULATION Ligands: ADP |
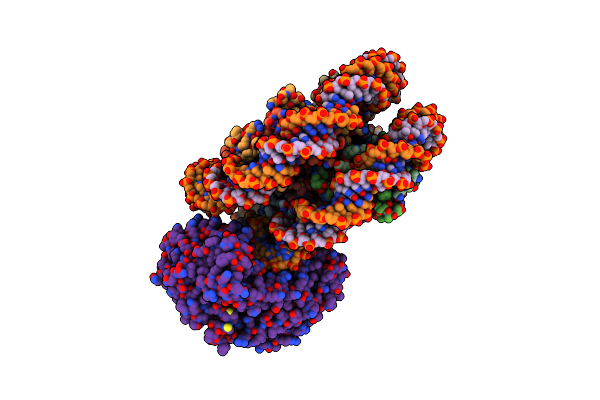 |
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: GENE REGULATION Ligands: BEF, ADP |
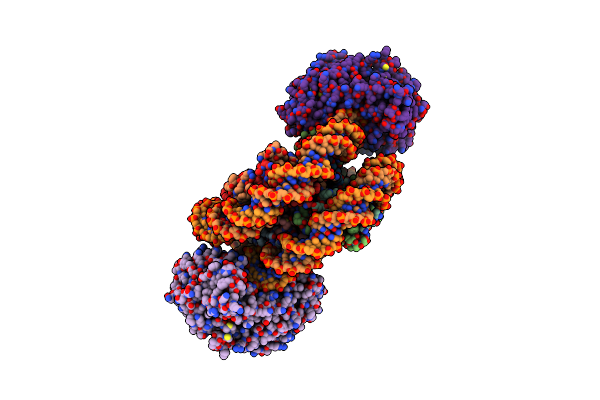 |
Structure Of Ddm1-Nucleosome Complex In The Adp-Befx State With Ddm1 Bound To Shl2 And Shl-2
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: GENE REGULATION Ligands: BEF, ADP |
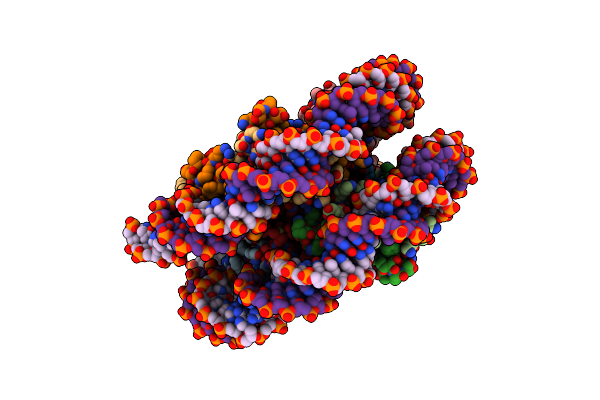 |
Organism: Arabidopsis thaliana, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: GENE REGULATION |
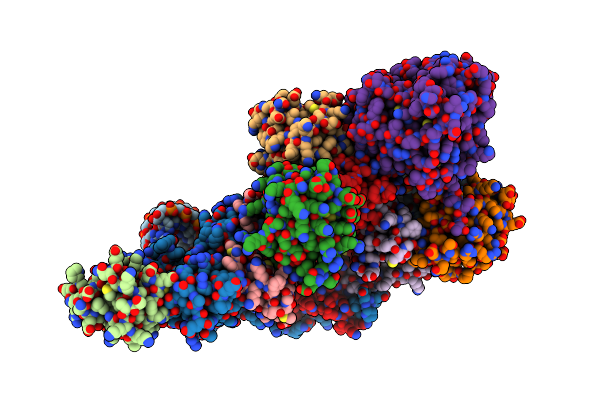 |
The H/Aca Rnp Lobe Of Human Telomerase With The Dyskerin Thumb Loop In A Semi-Closed Conformation
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2024-01-31 Classification: RNA BINDING PROTEIN |
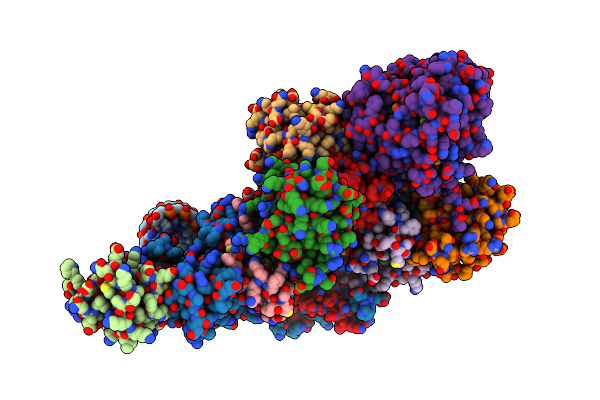 |
The H/Aca Rnp Lobe Of Human Telomerase With The Dyskerin Thumb Loop In An Open Conformation
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2024-01-31 Classification: RNA BINDING PROTEIN |
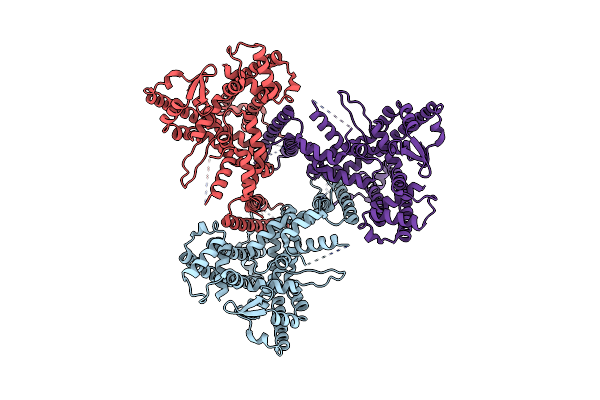 |
Structure Of Tissue-Specific Lipid Scramblase Atg9B Homotrimer, Refined With C3 Symmetry Applied
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-11-15 Classification: LIPID TRANSPORT |
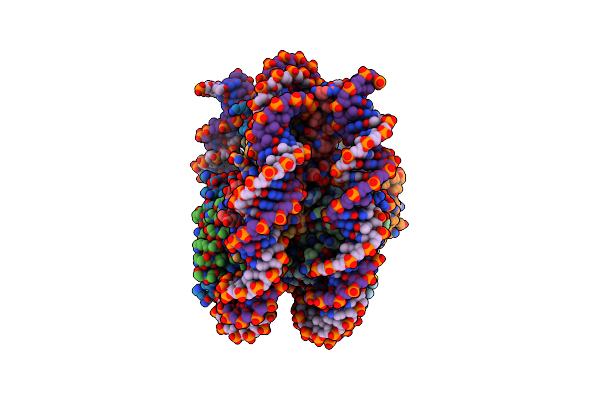 |
Arabidopsis Ddm1 Bound To Nucleosome (H2A.W, H2B, H3.3, H4, With 147 Bp Dna)
Organism: Arabidopsis thaliana, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-08-09 Classification: GENE REGULATION |
 |
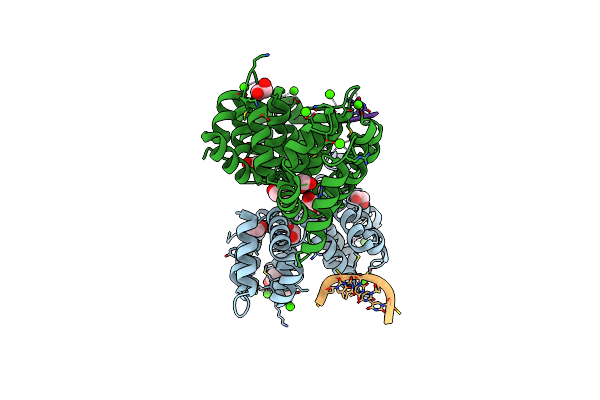
Crystal Structure Of Human Annexin A2 In Complex With Full Phosphorothioate 5-10 2'-Methoxyethyl Dna Gapmer Antisense Oligonucleotide Solved At 1.87 A Resolution
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2022-09-14 Classification: LIPID BINDING PROTEIN Ligands: GOL, CA, MG |
 |
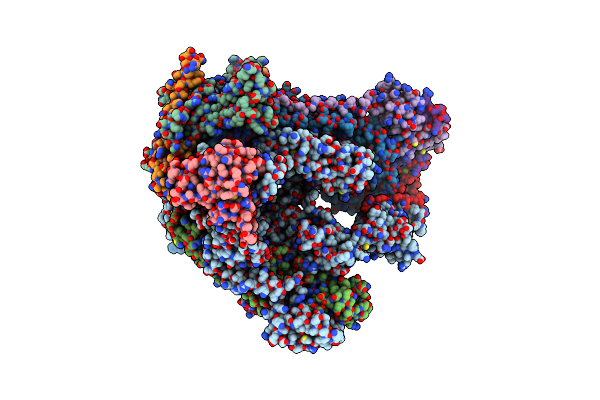
Crystal Structure Of Human Annexin A2 In Complex With Full Phosphorothioate 5-10 2'-Methoxyethyl Dna Gapmer Antisense Oligonucleotide Solved At 2.4 A Resolution
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-09-14 Classification: LIPID BINDING PROTEIN Ligands: CA, EDO |
 |
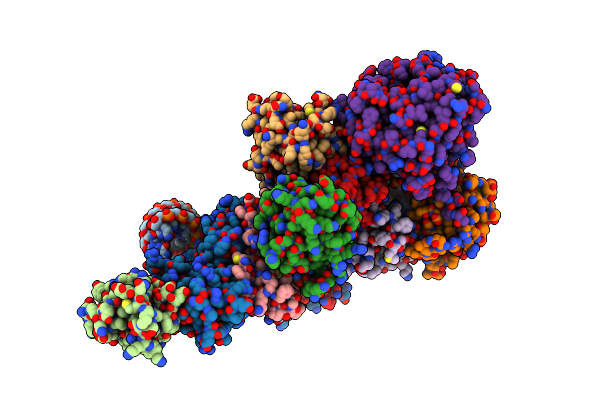
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2022-08-31 Classification: TRANSCRIPTION |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2022-04-20 Classification: REPLICATION |

