Search Count: 69
All
Selected
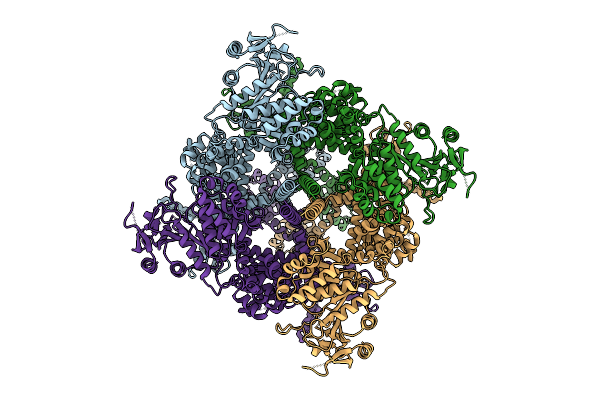 |
Avian Trpm8 (Parus Major) Desensitized, Fully-Swapped, Ligand-Free Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
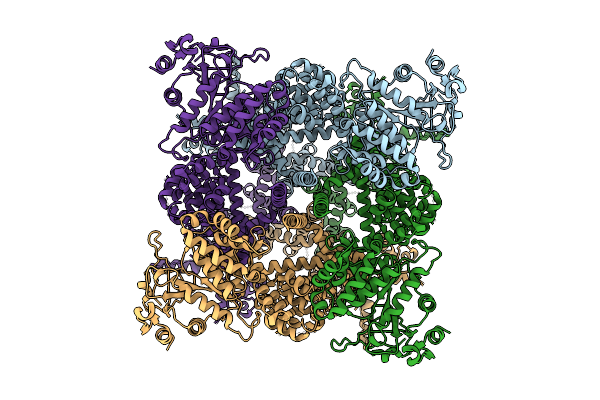 |
Avian Trpm8 (Parus Major) Menthol Bound Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: XUQ |
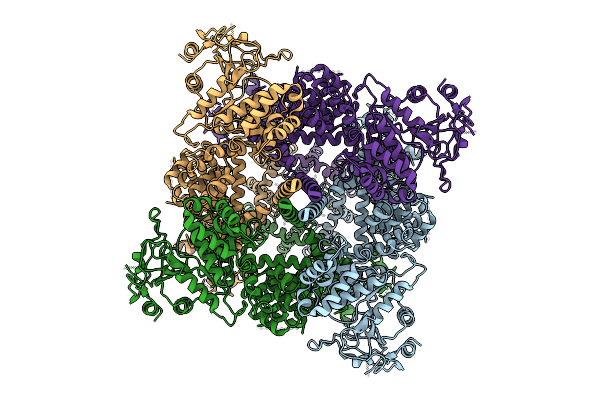 |
Avian Trpm8 (Parus Major) Semi-Swapped, Calcium Free, Menthol Bound Structure Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: XUQ |
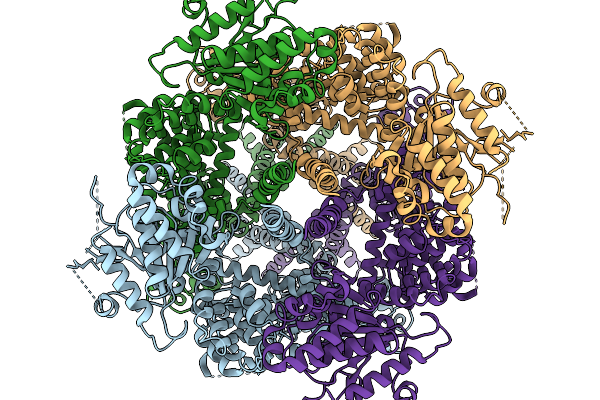 |
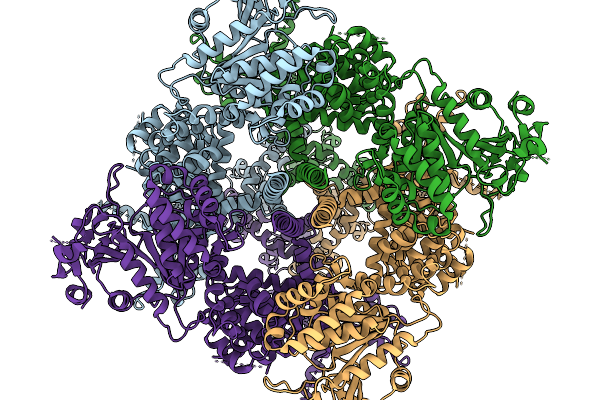
Avian Trpm8 (Parus Major) Closed, Ligand-Free Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
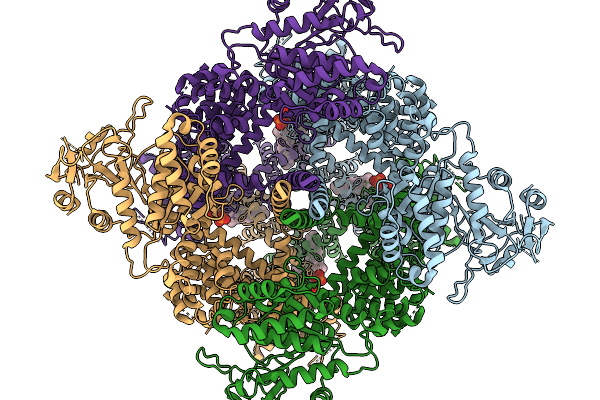
Avian Trpm8 (Parus Major) Semi-Swapped, Ligand-Free Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Avian Trpm8 (Parus Major) Semi-Swapped, Ligand-Free Structure At High Ph And 4 Degrees Celsius Resolved In Gdn Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: PT5 |
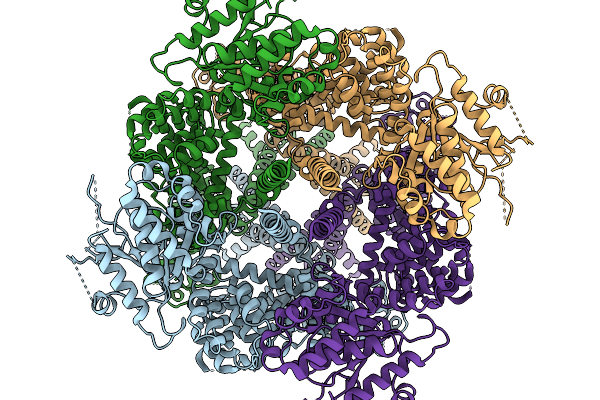 |
Avian Trpm8 (Parus Major) Fully-Swapped, Closed, Ligand-Free In The Presence Of Calcium, Structure Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
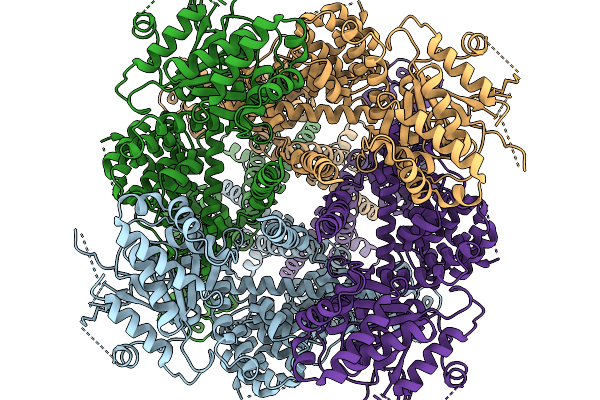
Avian Trpm8 (Parus Major) Semi-Swapped, Closed, Ligand-Free In The Presence Of Calcium, Structure Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: CA |
 |
Avian Trpm8 (Parus Major) Fully Swapped Closed, Calcium Free, In The Presence Of Menthol, Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN |
 |
Avian Trpm8 (Parus Major) Semi-Swapped, Closed, Calcium Free, Menthol Bound Structure Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: XUQ |
 |
Parus Major Trpm8 With A Chimeric Human Outer Pore Loops, Semi-Swapped, Ligand-Free, Cold, Structure Resolved In Gdn
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
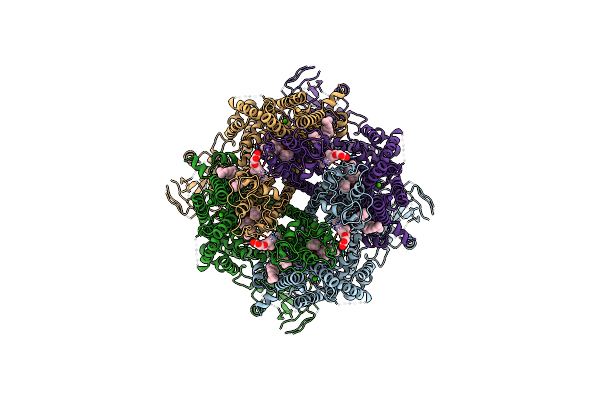 |
Cryo-Em Structure Of The Avian Great Tit Trpm8 Channel In Complex With The Antagonist Tc-I 2014
Organism: Parus major
Method: ELECTRON MICROSCOPY Resolution:3.26 Å Release Date: 2024-08-21 Classification: MEMBRANE PROTEIN Ligands: CA, Y01, T14 |
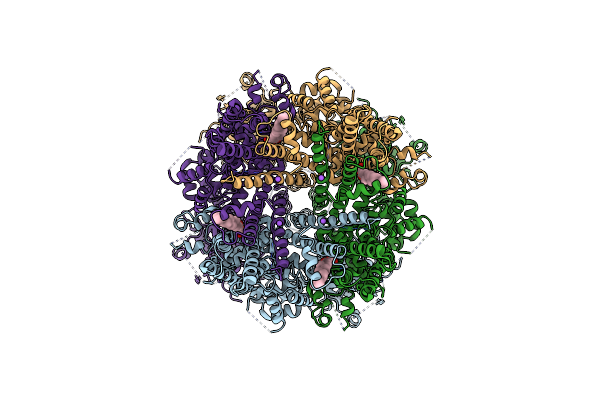 |
Structure Of The Trpm8 Cold Receptor By Single Particle Electron Cryo-Microscopy, Ligand-Free State
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2019-09-18 Classification: TRANSPORT PROTEIN Ligands: Y01, NA |
 |
Structure Of The Trpm8 Cold Receptor By Single Particle Electron Cryo-Microscopy, Amtb-Bound State
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2019-09-18 Classification: TRANSPORT PROTEIN Ligands: Y01, UND, 9PE, LQ7, NA |
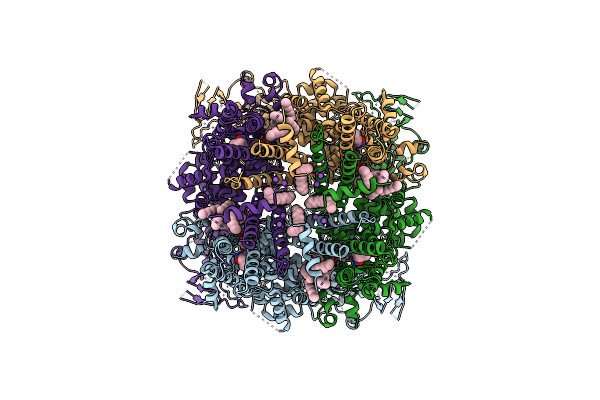 |
Structure Of The Trpm8 Cold Receptor By Single Particle Electron Cryo-Microscopy, Tc-I 2014-Bound State
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2019-09-18 Classification: TRANSPORT PROTEIN Ligands: Y01, UND, 9PE, T14, NA |
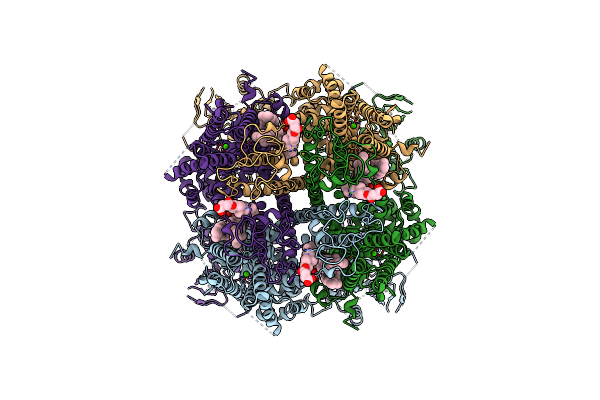 |
Structure Of The Trpm8 Cold Receptor By Single Particle Electron Cryo-Microscopy, Calcium-Bound State
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2019-09-18 Classification: TRANSPORT PROTEIN Ligands: CA, Y01 |
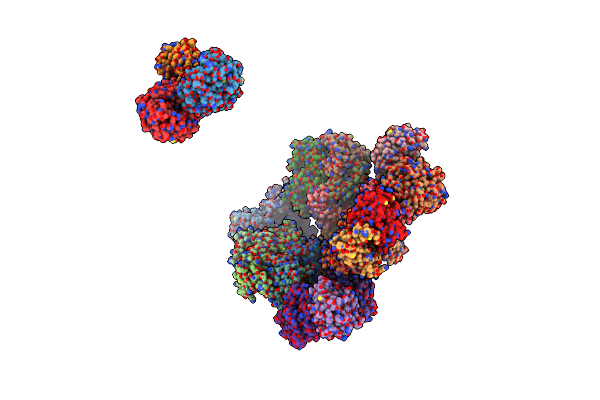 |
Crystal Structure Of Designed Two-Component Self-Assembling Icosahedral Cage I53-40
Organism: Methanocaldococcus jannaschii (strain atcc 43067 / dsm 2661 / jal-1 / jcm 10045 / nbrc 100440), Vibrionales bacterium (strain swat-3)
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2016-07-27 Classification: PROTEIN BINDING |
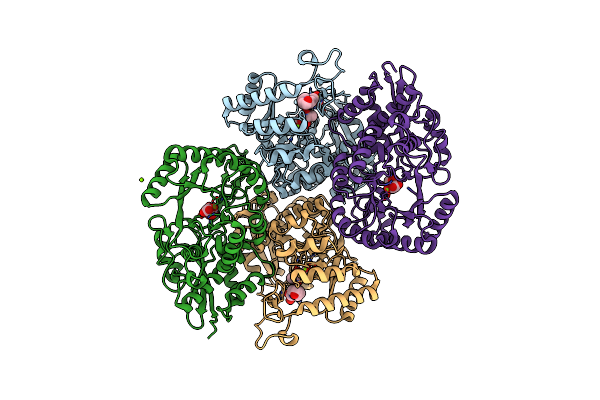 |
Crystal Structure Of The Mutant P311A Of Enolase Superfamily Member From Vibrionales Bacterium Complexed With Mg And D-Arabinonate
Organism: Vibrionales bacterium swat-3
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2012-06-06 Classification: ISOMERASE Ligands: MG, EPE, D8T |
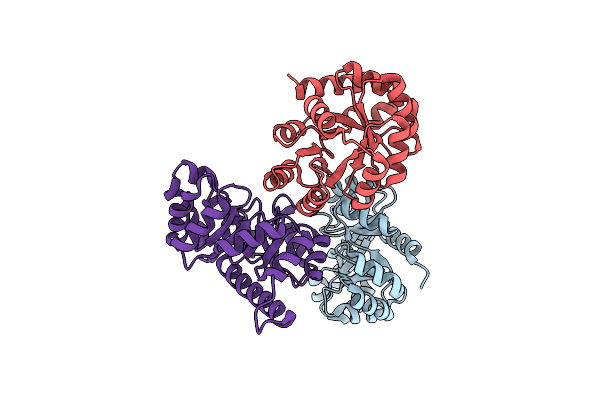 |
Crystal Structure Of Probable Keto-Hydroxyglutarate-Aldolase From Vibrionales Bacterium Swat-3 (Target Efi-502156)
Organism: Vibrionales bacterium swat-3
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2012-03-21 Classification: LYASE Ligands: CL |
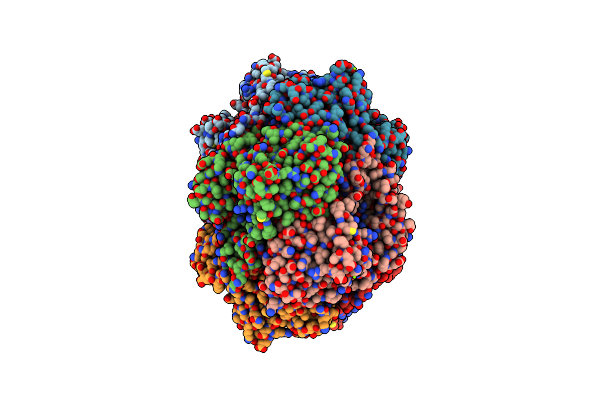 |
Crystal Structure Of Enolase Superfamily Member From Vibrionales Bacterium Complexed With Mg And Glycerol In The Active Site
Organism: Vibrionales bacterium swat-3
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2012-03-14 Classification: ISOMERASE Ligands: MG, GOL |

