Search Count: 9,001
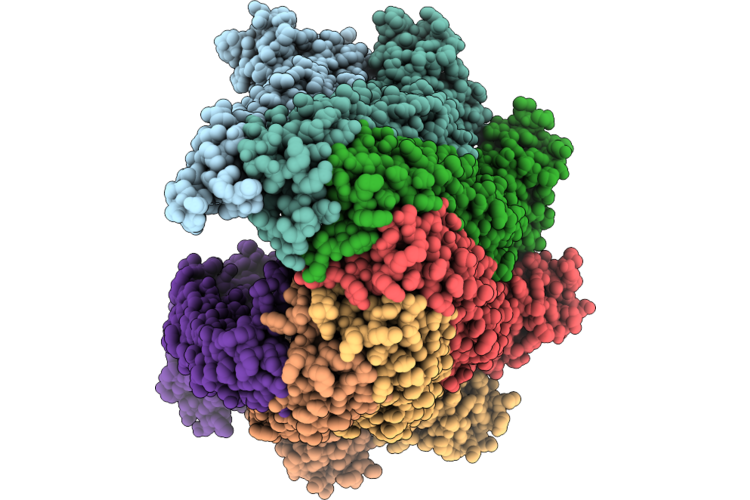 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
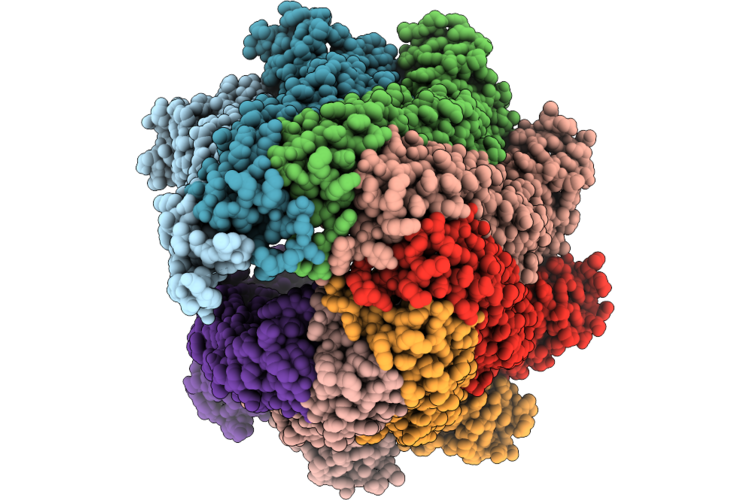 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
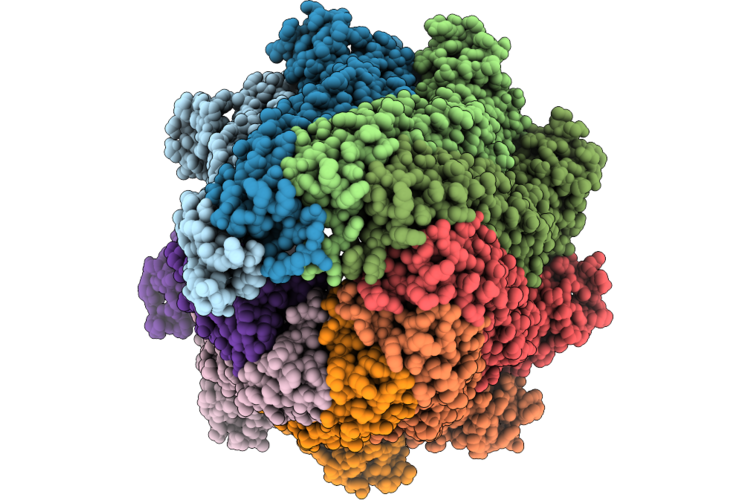 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
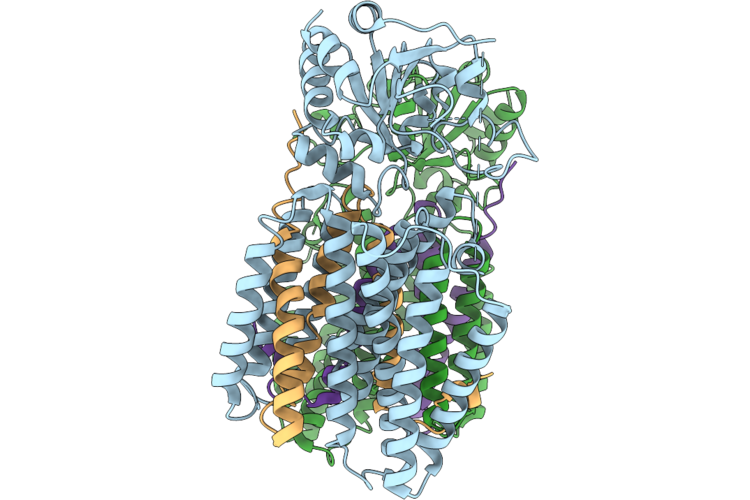 |
Single-Subunit Apo Escherichia Coli Nicotinamide Nucleotide Transhydrogenase
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
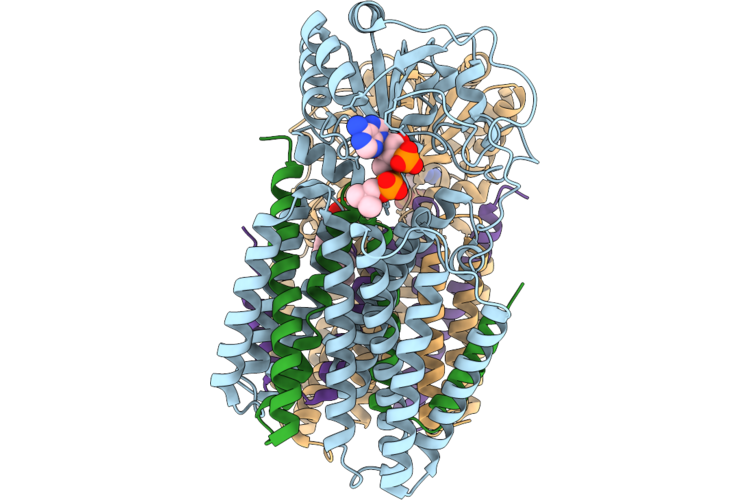 |
Single-Subunit Escherichia Coli Nicotinamide Nucleotide Transhydrogenase In The Presence Of Palmitoyl-Coa
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: COA, PC1 |
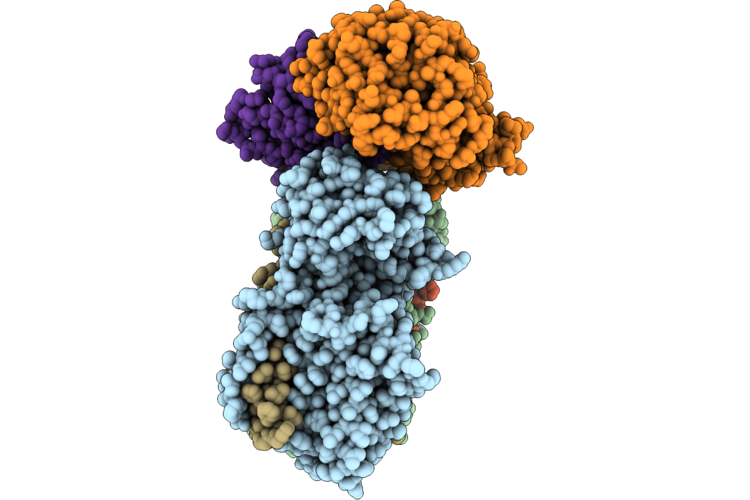 |
Single-Subunit Apo Escherichia Coli Nicotinamide Nucleotide Transhydrogenase
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
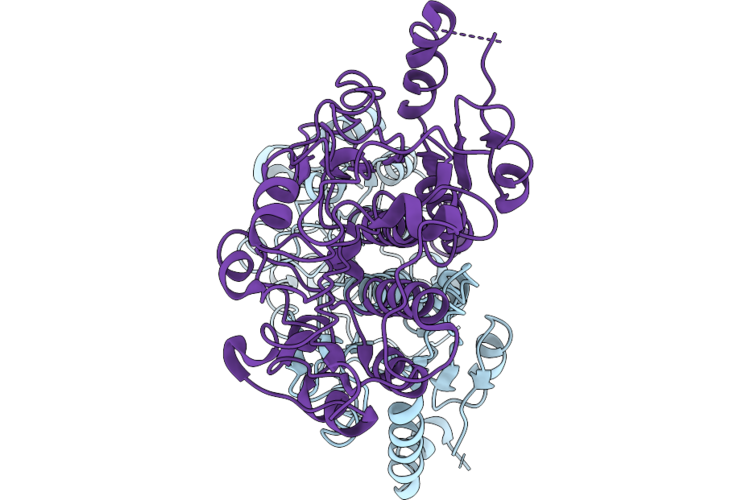
Structures Of Multiple States Of The Nicotinamide Nucleotide Transhydrogenase From Escherichia Coli
Organism: Escherichia coli (strain k12)
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
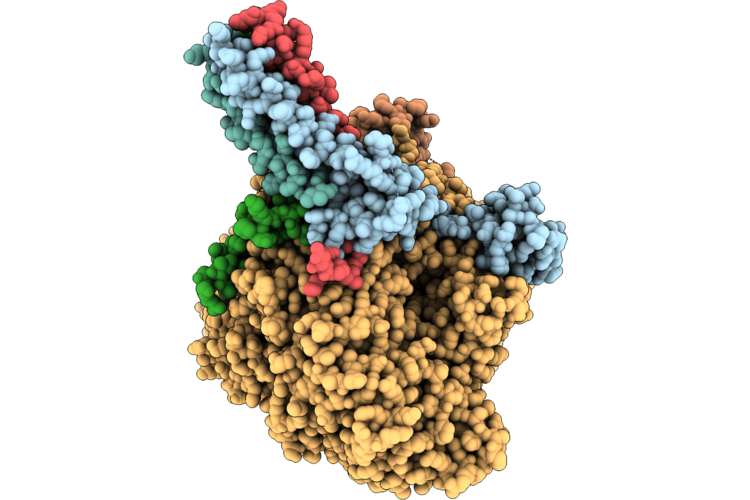
Escherichia Coli Transhydrogenase The Dissociated (Di)2 Dimer In The Presence Of Both Nadph And Nadp+
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of The Human Measles Virus Rna-Dependent Rna Polymerase Complex Bound To Viral Protein C
Organism: Escherichia coli k-12, Measles virus genotype a-vaccine, Synthetic construct, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: ZN |
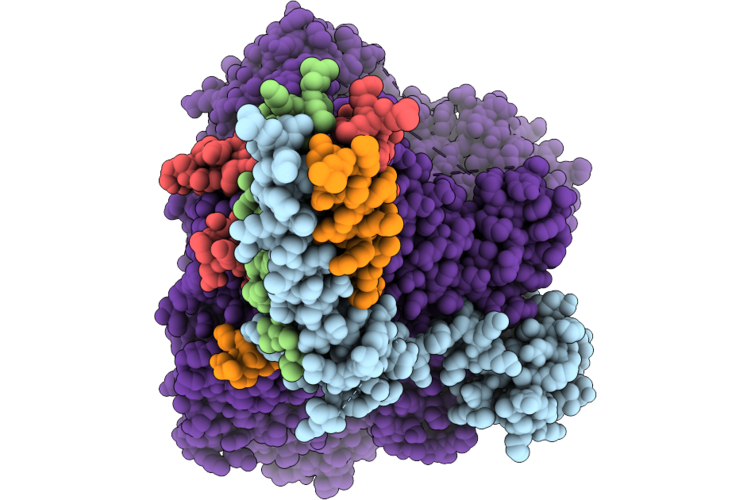 |
Organism: Escherichia coli k-12, Measles virus genotype a-vaccine, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: ZN |
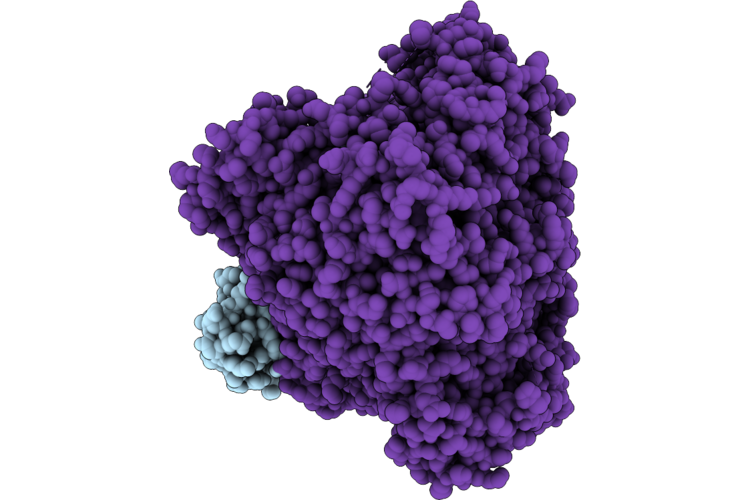 |
Cryo-Em Structure Of The Human Measles Virus Rna-Dependent Rna Polymerase Complex
Organism: Escherichia coli k-12, Measles virus genotype a-vaccine, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: ZN |
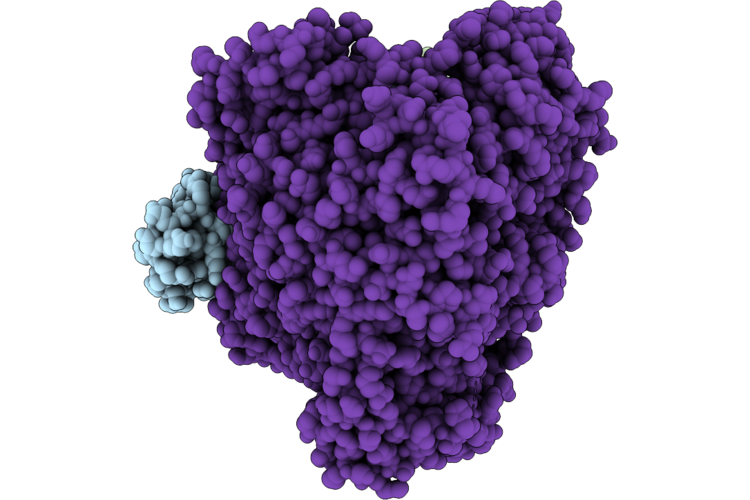 |
Cryo-Em Structure Of The Human Measles Virus Rna-Dependent Rna Polymerase Bound To Allosteric Inhibitor Erdrp-0519
Organism: Escherichia coli k-12, Measles virus genotype a-vaccine, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: ZN, A1EF9 |
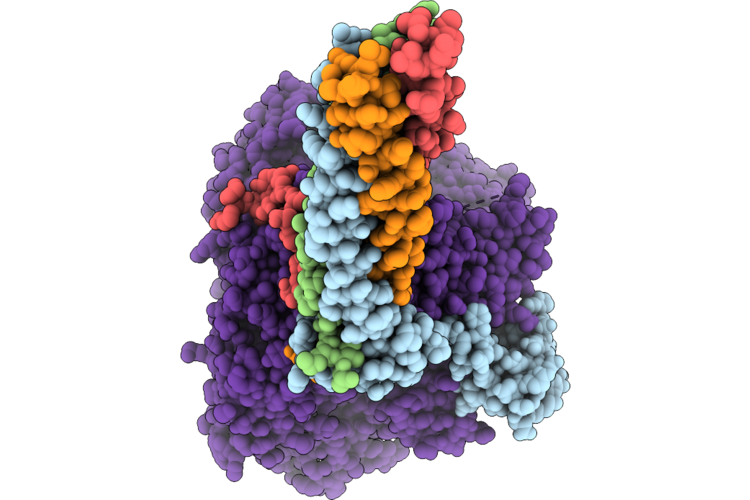 |
Cryo-Em Structure Of The Nipah Virus Rna-Dependent Rna Polymerase Complex Bound To Allosteric Inhibitor Erdrp-0519
Organism: Escherichia coli k-12, Henipavirus nipahense
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-27 Classification: VIRAL PROTEIN Ligands: ZN, A1EF9 |
 |
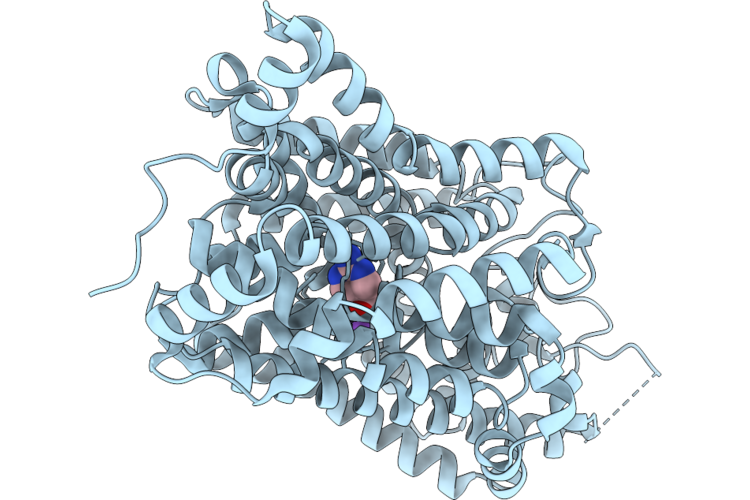
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: A1EVN, CL |
 |
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: A1EMK, CL, NA |
 |
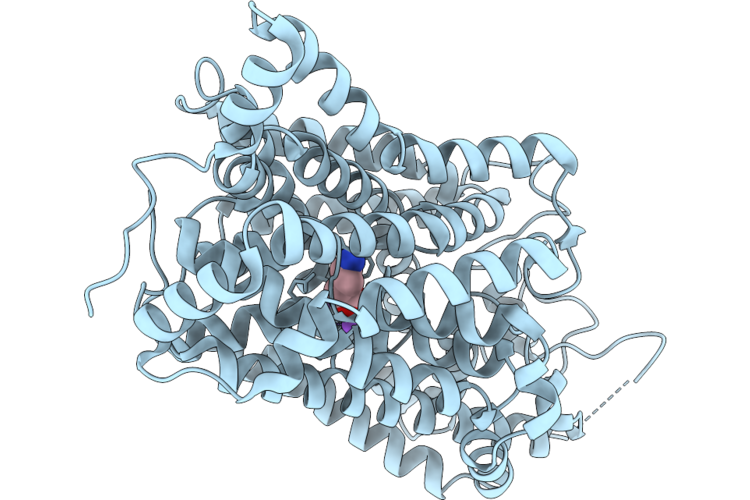
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: BET, NA, CL |
 |
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: ABU, CL, NA |
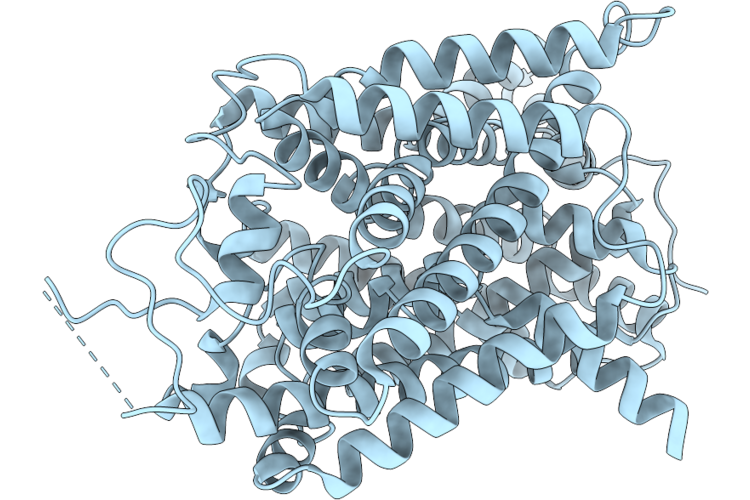 |
Structure Of The Apo State Of Human Betaine/Gaba Transporter 1 In The Inward-Facing Conformation
Organism: Homo sapiens, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-05-27 Classification: MEMBRANE PROTEIN Ligands: CL |

