Search Count: 1,677
All
Selected
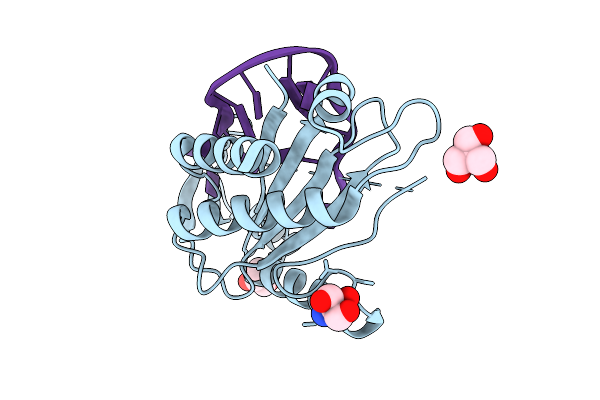 |
Crystal Structure Of The Catalytic N-Terminal Domain Of The Relaxase Of Plasmid Pls20 In Complex With The Core Region
Organism: Bacillus subtilis subsp. natto
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: TRS |
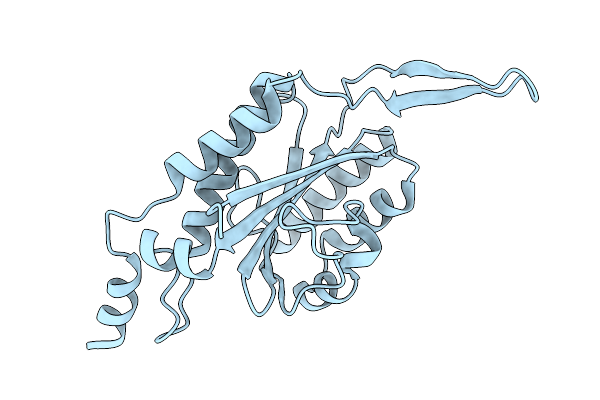 |
Crystal Structure Of The Relaxase Domain Of Relpls20, Prototype Of Mobl Family Of Relaxases
Organism: Bacillus subtilis subsp. natto
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: CL |
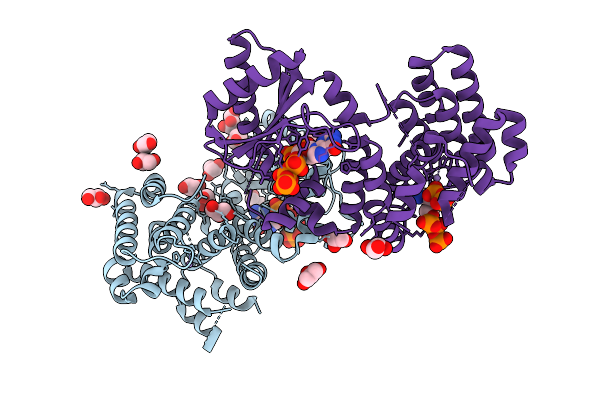 |
Crystal Structure Of The Relseq N-Terminal Domain From Streptococcus Equisimilis In Complex With Pppgpp
Organism: Streptococcus dysgalactiae subsp. equisimilis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: GOL, MN, 0O2, CL |
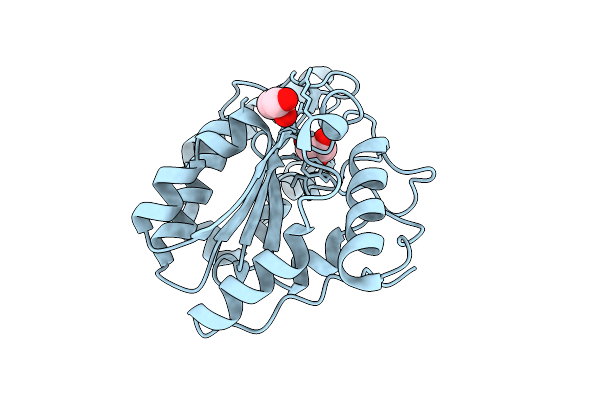 |
Organism: Micrococcus flavus
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN Ligands: GOL |
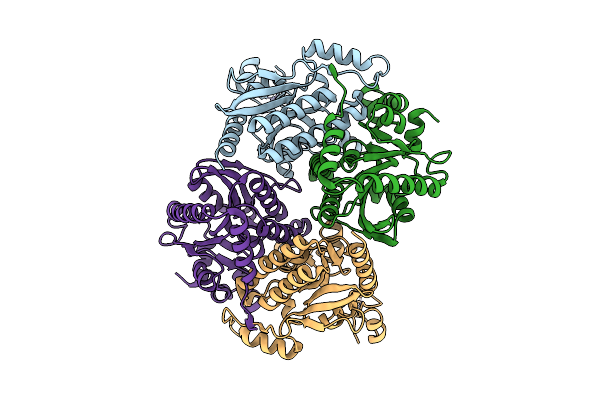 |
Crystal Structure Of The Carboxyltransferase Subunit Of Acetyl-Coa Carboxylase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2026-03-11 Classification: LIGASE Ligands: ZN |
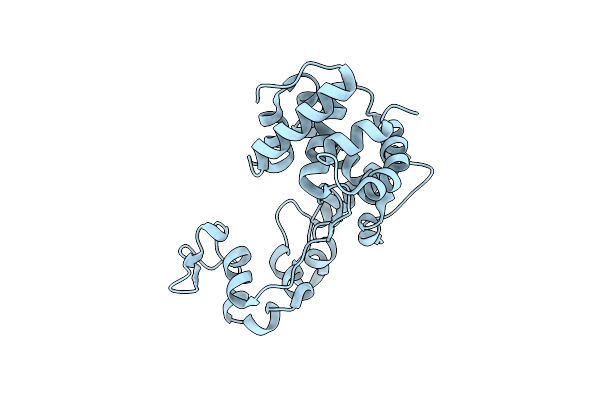 |
Organism: Aeromonas dhakensis
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2025-04-23 Classification: HYDROLASE |
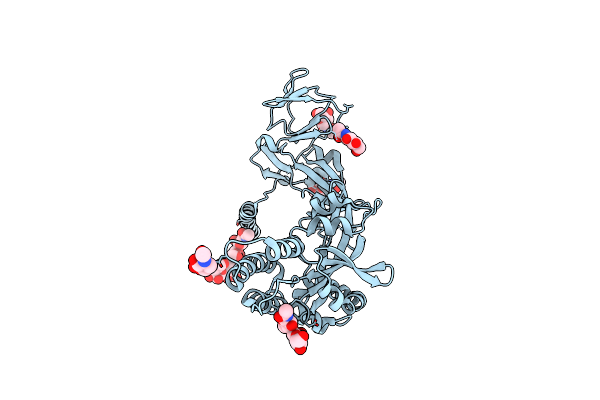 |
Organism: Mumps orthorubulavirus
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2025-02-19 Classification: VIRAL PROTEIN Ligands: NAG |
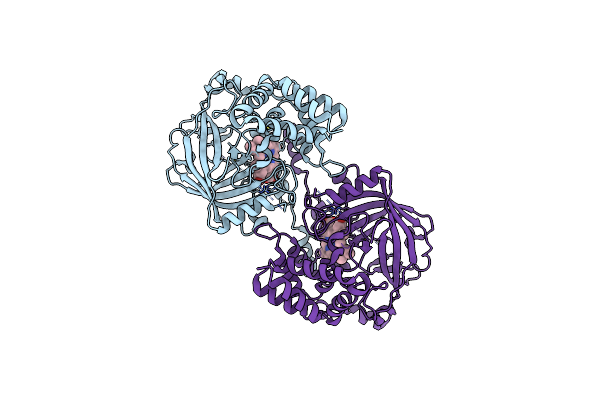 |
Organism: Claviceps fusiformis
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: OXIDOREDUCTASE Ligands: HEM |
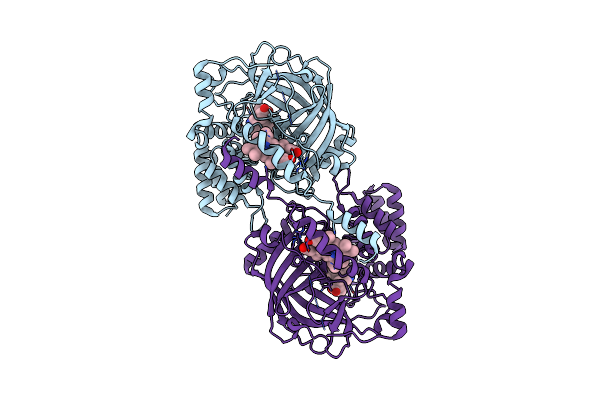 |
Structure Of Chanoclavine Synthase From Claviceps Fusiformis In Complex With Prechanoclavine
Organism: Claviceps fusiformis
Method: ELECTRON MICROSCOPY Resolution:2.33 Å Release Date: 2025-01-01 Classification: OXIDOREDUCTASE Ligands: HEM, LH6 |
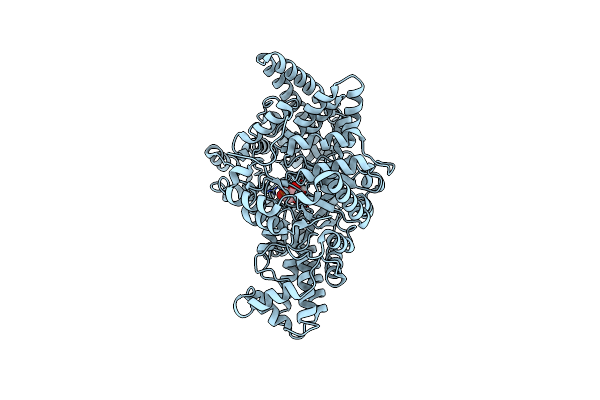 |
Crystal Structure Of Pfld Bound To 1,5-Anhydromannitol-6-Phosphate In Streptococcus Dysgalactiae Subsp. Equisimilis
Organism: Streptococcus dysgalactiae subsp. equisimilis
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-10-02 Classification: LYASE Ligands: 7P6 |
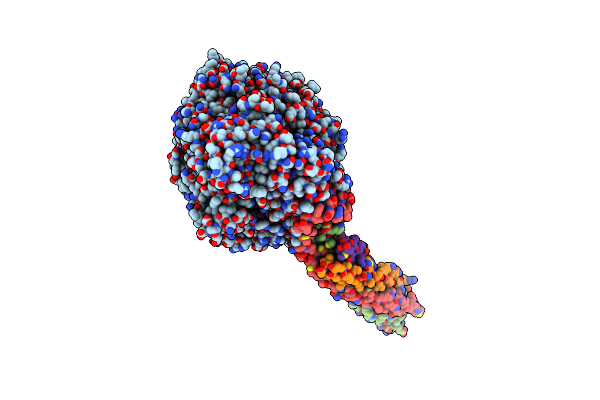 |
Structure Of The Mumps Virus L Protein (State2) Bound By Phosphoprotein Tetramer
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN |
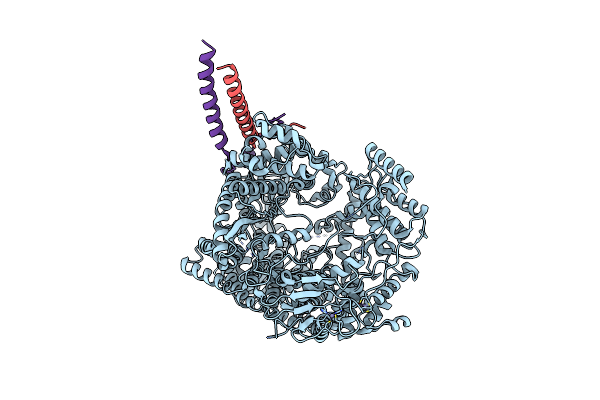 |
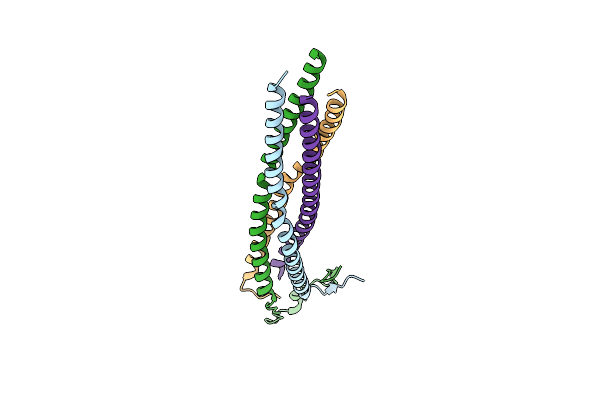
Structure Of N-Terminal Domain Of L Protein Bound With Phosphoprotein From Mumps Virus
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN |
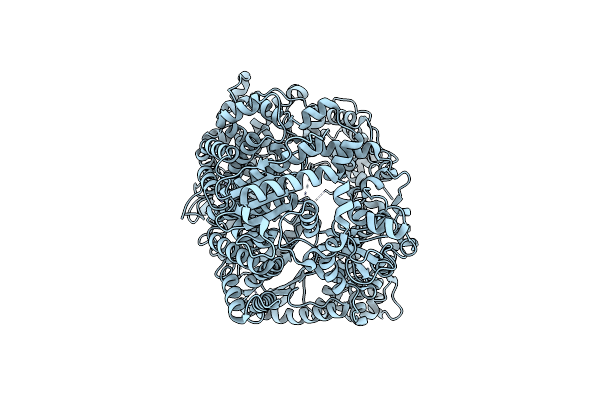 |
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Organism: Mumps orthorubulavirus
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: VIRAL PROTEIN |
 |
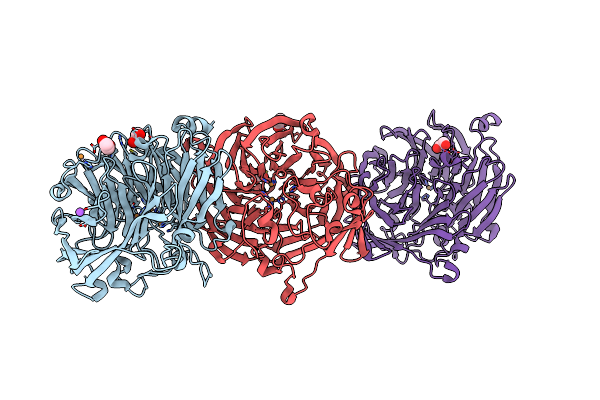
The Structure Of Non-Activated Thiocyanate Dehydrogenase From Pelomicrobium Methylotrophicum (Pmtcdh)
Organism: Pelomicrobium methylotrophicum
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-05-08 Classification: OXIDOREDUCTASE Ligands: EDO, CL, CU, PEG |
 |
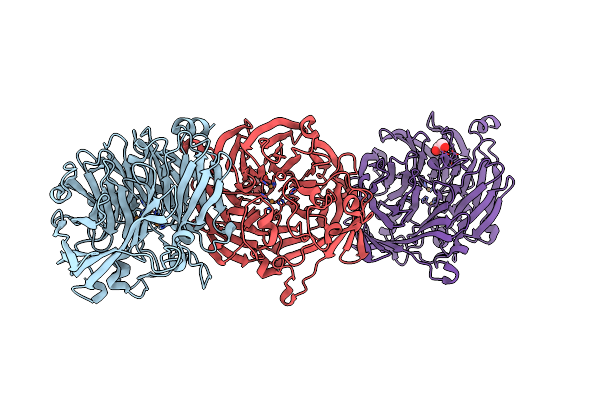
The Structure Of Thiocyanate Dehydrogenase From Pelomicrobium Methylotrophicum (Pmtcdh), Activated By Crystals Soaking With 1 Mm Cucl2 During 6 Months
Organism: Pelomicrobium methylotrophicum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-05-08 Classification: OXIDOREDUCTASE Ligands: EDO, CU, NA |
 |
The Structure Of Thiocyanate Dehydrogenase From Pelomicrobium Methylotrophicum (Pmtcdh), Activated By Crystals Soaking With 1 Mm Cucl2 And Na Ascorbate During 12 Hours
Organism: Pelomicrobium methylotrophicum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-05-08 Classification: OXIDOREDUCTASE Ligands: CU, EDO |
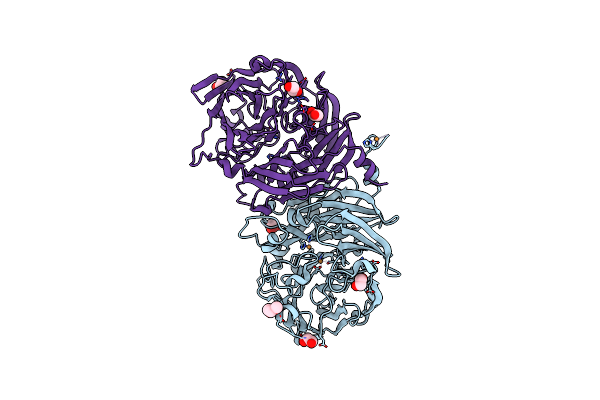 |
The Structure Of Non-Activated Thiocyanate Dehydrogenase Mutant With The H447Q Substitution From Pelomicrobium Methylotrophicum (Pmtcdh H447Q)
Organism: Pelomicrobium methylotrophicum
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-04-24 Classification: OXIDOREDUCTASE Ligands: PEG, EDO, CU, CL |

