Search Count: 3,123
All
Selected
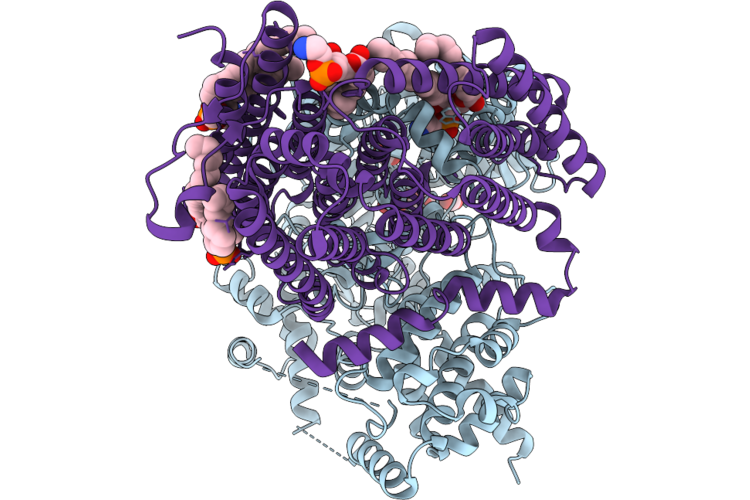 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
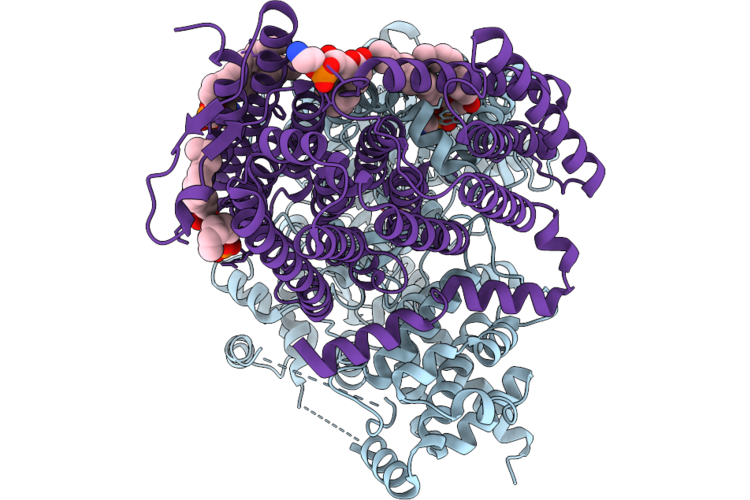 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, BCT, CO2, 6OU |
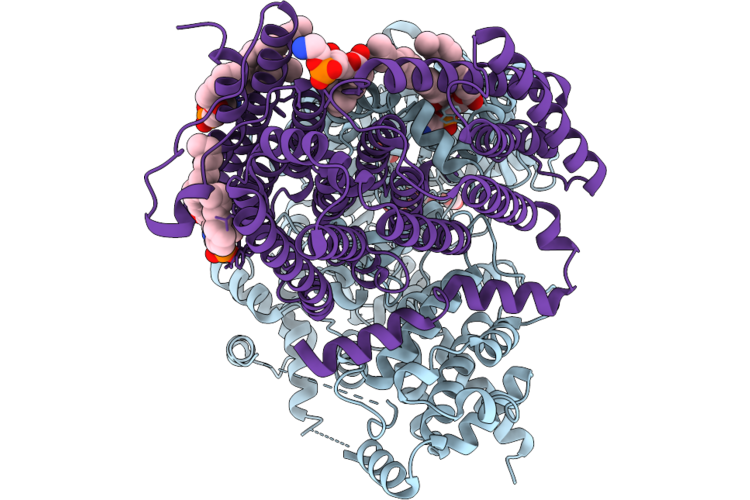 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
 |
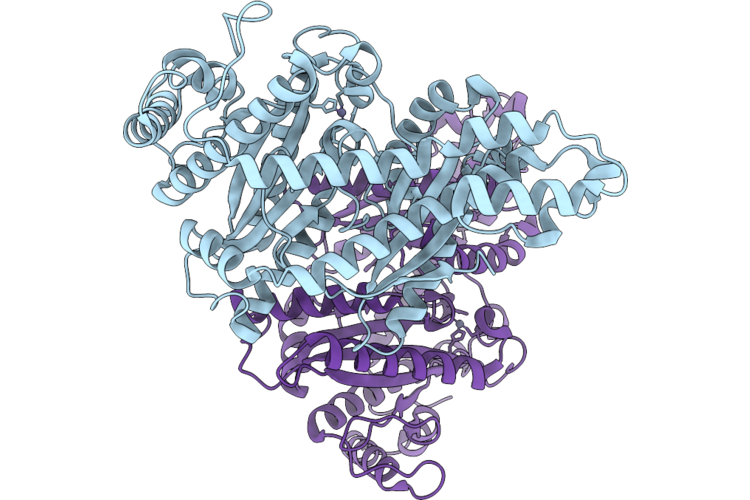
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
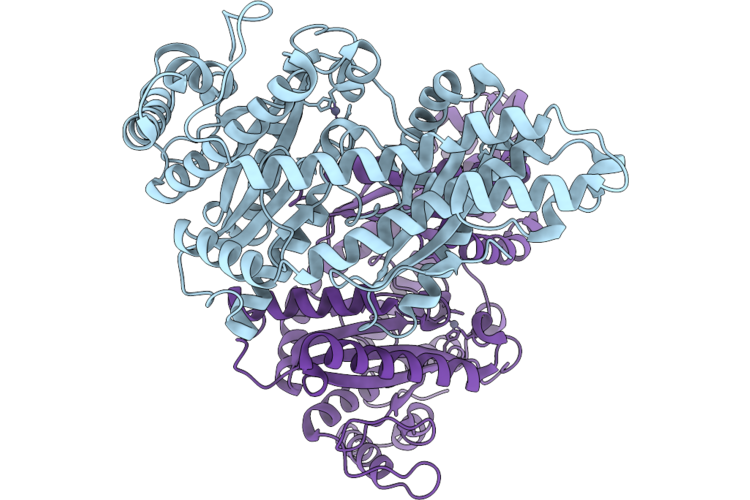 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 1
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 2
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.22 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of Self-Assembled Zymomonas Mobilis Levansucrase Nanotube
Organism: Zymomonas mobilis subsp. mobilis atcc 10988
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSFERASE |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv37A (Compound 9)
Organism: Mycobacterium tuberculosis '98-r604 inh-rif-em'
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG0 |
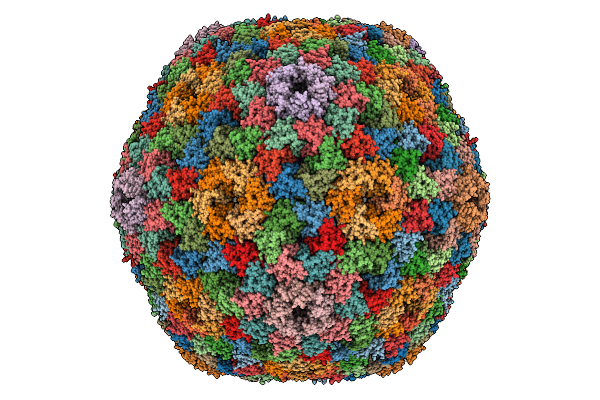 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
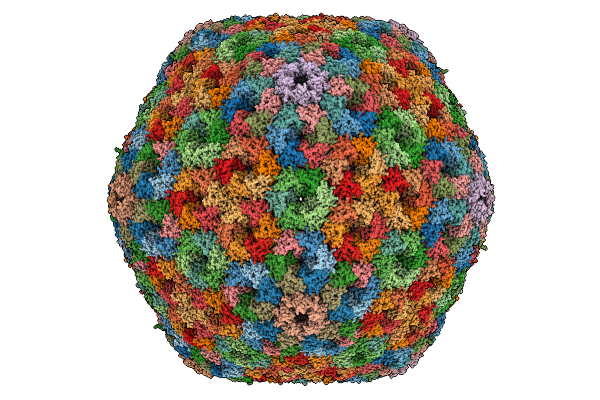 |
Cryo-Em Structure Of Carboxysomal Mid-Shell: T = 16 Shell Under C1 Symmetry.
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
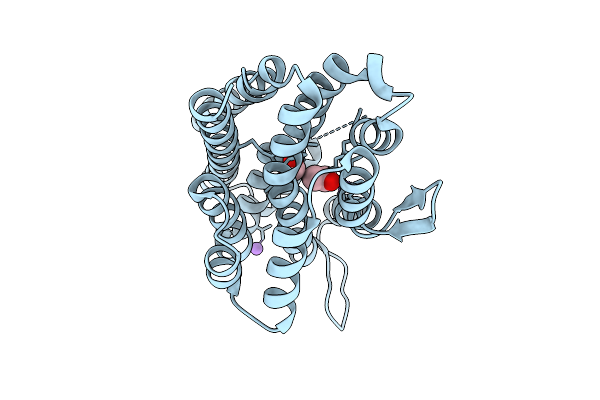 |
Organism: Burkholderia cenocepacia h111
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-11-12 Classification: METAL BINDING PROTEIN Ligands: FE, ACY, FMT, NA |
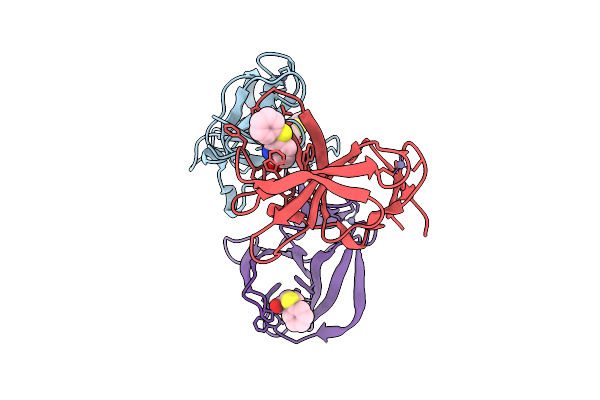 |
Cereblon Isoform 4 From Magnetospirillum Gryphiswaldense In Complex With Glutarimide Based Compound 2N
Organism: Magnetospirillum gryphiswaldense
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: ZN, A1IXA, A1I4R |
 |
Cereblon Isoform 4 From Magnetospirillum Gryphiswaldense In Complex With Glutarimide Based Compound 2J
Organism: Magnetospirillum gryphiswaldense
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: ZN, A1IW9, PO4 |
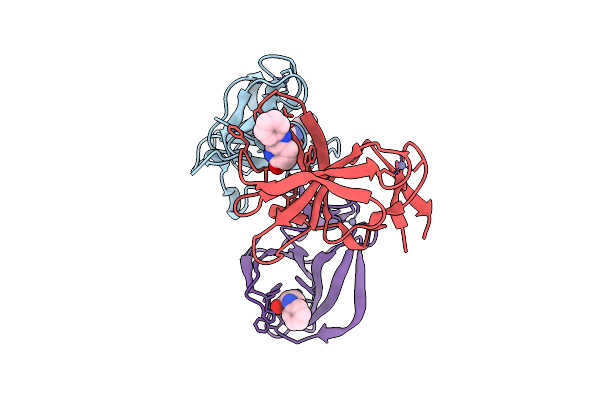 |
Cereblon Isoform 4 From Magnetospirillum Gryphiswaldense In Complex With Glutarimide Based Compound 2O
Organism: Magnetospirillum gryphiswaldense
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: ZN, A1IW8, A1I4P |

