Search Count: 2,673
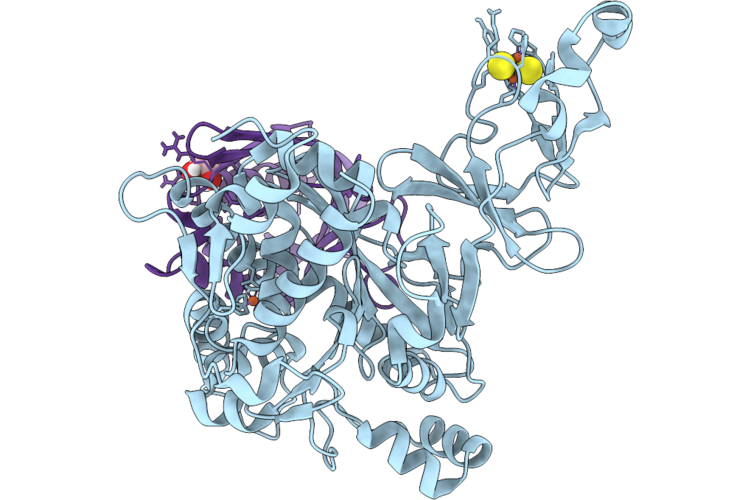 |
Organism: Variovorax paradoxus
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2026-05-20 Classification: METAL BINDING PROTEIN Ligands: FES, FE2, EDO |
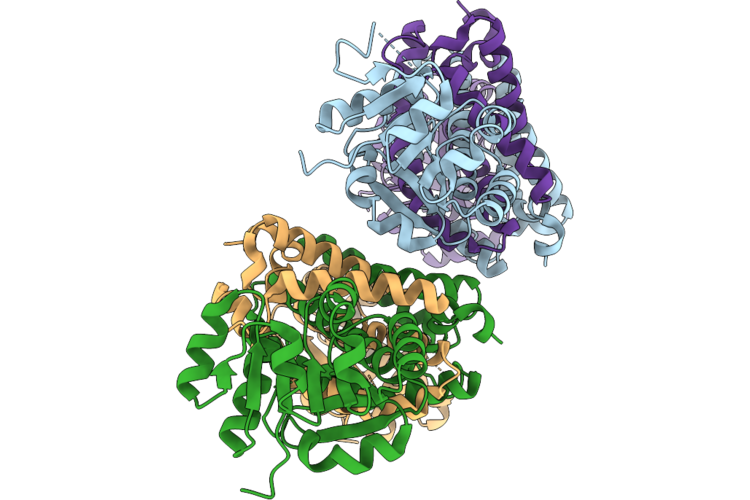 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE |
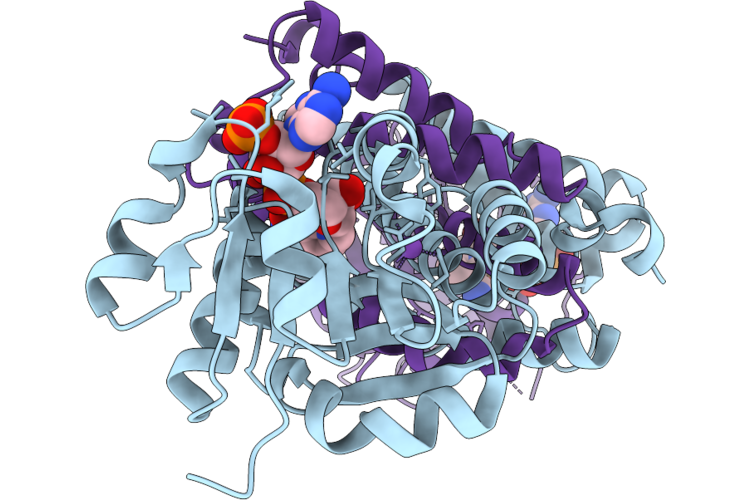 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: NAP, NA |
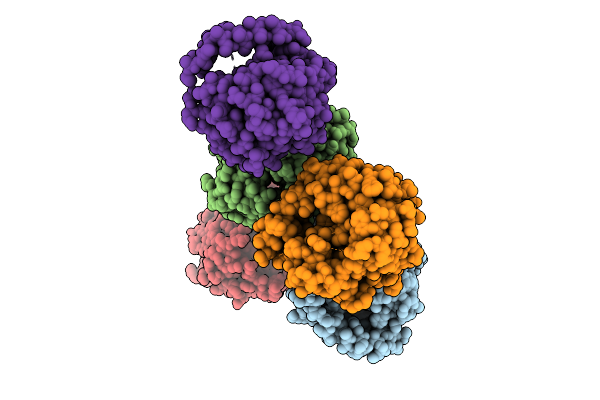 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Faecium (S126A Mutant)
Organism: Sphingobacterium faecium
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2026-04-08 Classification: HYDROLASE |
 |
Organism: Salmonella enterica subsp. enterica serovar bredeney
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: ANTIVIRAL PROTEIN |
 |
Organism: Homo sapiens, Sars bat coronavirus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: RIBOSOME |
 |
Organism: Homo sapiens, Sars bat coronavirus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: RIBOSOME |
 |
Organism: Homo sapiens, Sars bat coronavirus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: RIBOSOME |
 |
Organism: Homo sapiens, Sars bat coronavirus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: RIBOSOME |
 |
Organism: Homo sapiens, Sars bat coronavirus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: RIBOSOME |
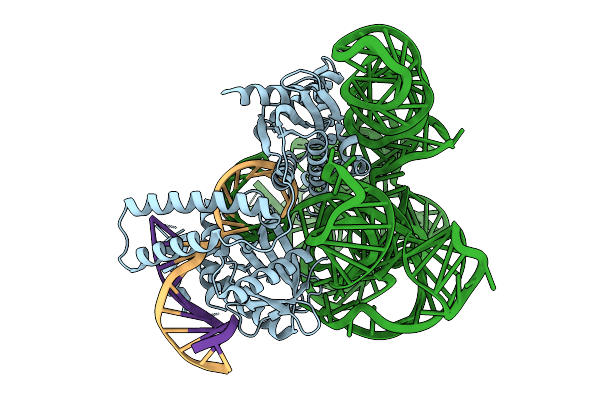 |
Organism: Scytonema hofmannii, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-09-17 Classification: DNA BINDING PROTEIN |
 |
(1-Methylalkyl)Succinate Synthase Alpha-Beta-Gamma-Delta Complex With Bound Fumarate
Organism: Azoarcus sp. hxn1
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: LYASE Ligands: FUM, DTT, SF4, FE |
 |
Organism: Riemerella anatipestifer
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2025-06-25 Classification: TRANSPORT PROTEIN |
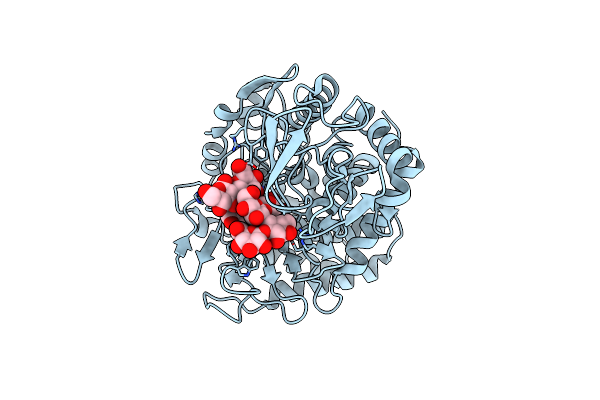 |
Structure Of Beta-1,2-Glucanase From Xanthomonas Campestris Pv. Campestris (Beta-1,2-Glucoheptasaccharide Complex)-E239Q Mutant
Organism: Xanthomonas campestris pv. campestris
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-01-22 Classification: HYDROLASE |
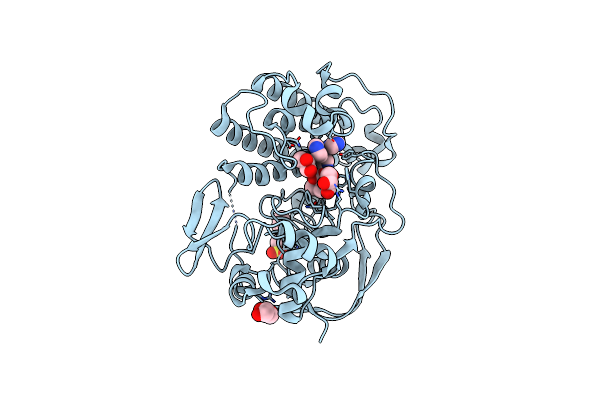 |
Organism: Variovorax paradoxus
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: NI, AVJ, IMD, GOL, NA, NHE |
 |
Organism: Variovorax paradoxus
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: UD1 |
 |
Conformational Landscape Of The Type V-K Crispr-Associated Transposonintegration Assembly Cast V-K Composite Map
Organism: Scytonema hofmannii
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: DNA BINDING PROTEIN Ligands: MG, ZN, ATP |
 |
Conformational Landscape Of The Type V-K Crispr-Associated Transposonintegration Assembly Cast V-K Cas12K Domain Local-Refinement Map
Organism: Scytonema hofmannii
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: DNA BINDING PROTEIN Ligands: MG, ZN |
 |
Conformational Landscape Of The Type V-K Crispr-Associated Transposonintegration Assembly Cast V-K Tnsc Domain Local-Refinement Map
Organism: Scytonema hofmannii
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: DNA BINDING PROTEIN Ligands: MG, ATP |
 |
Conformational Landscape Of The Type V-K Crispr-Associated Transposonintegration Assembly Cast V-K Tnsb Domain Local-Refinement Map
Organism: Scytonema hofmannii
Method: ELECTRON MICROSCOPY Release Date: 2024-06-19 Classification: DNA BINDING PROTEIN Ligands: MG |

