Search Count: 4
All
Selected
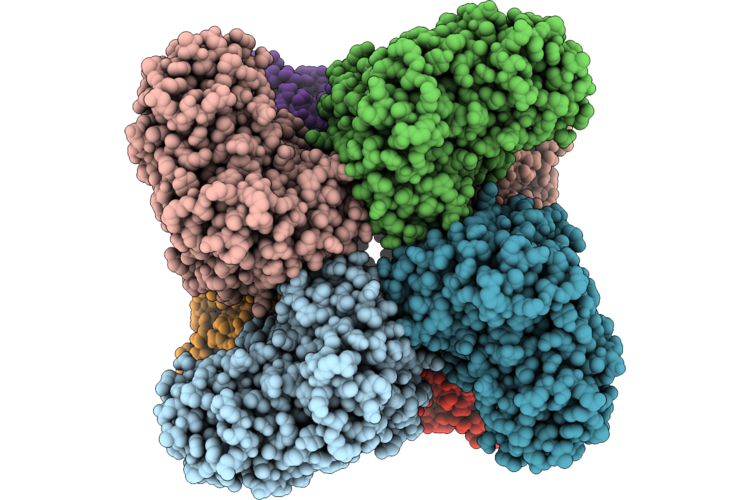 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
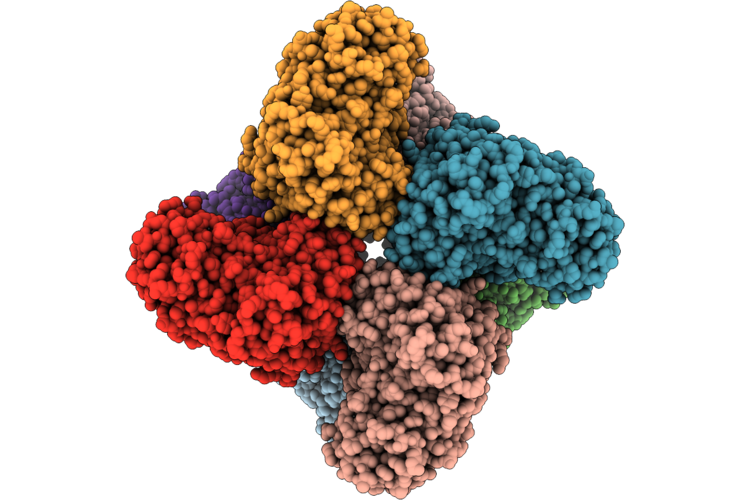 |
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
 |
Crystal Structure Of Monomeric Plp-Dependent Transaminase From Desulfobacula Toluolica In F 41 3 2 Space Group
Organism: Desulfobacula toluolica
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2024-09-18 Classification: TRANSFERASE |
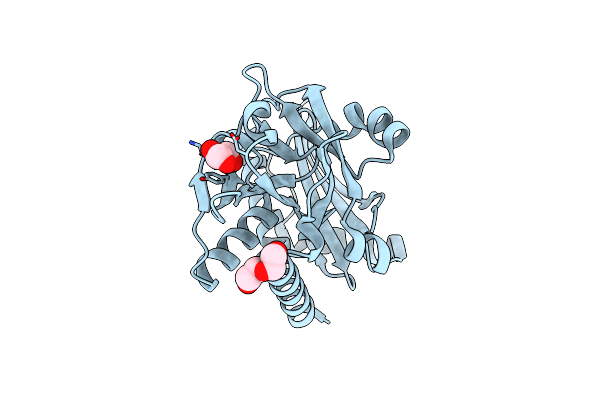 |
Crystal Structure Of Monomeric Plp-Dependent Transaminase From Desulfobacula Toluolica In P 21 21 21 Space Group
Organism: Desulfobacula toluolica
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2024-09-18 Classification: TRANSFERASE Ligands: PEG, GOL |

