Search Count: 1,147
All
Selected
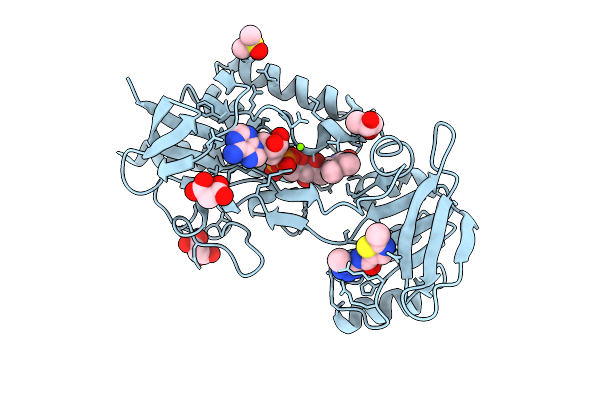 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2997505083
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1JAC, DMS |
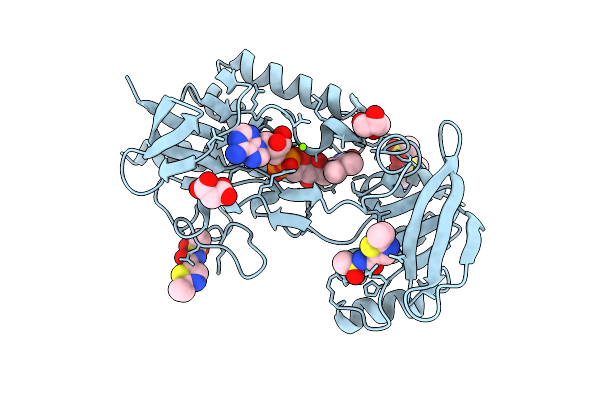 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z854627136
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I94 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z30008604
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I96 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2858787682
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I97 |
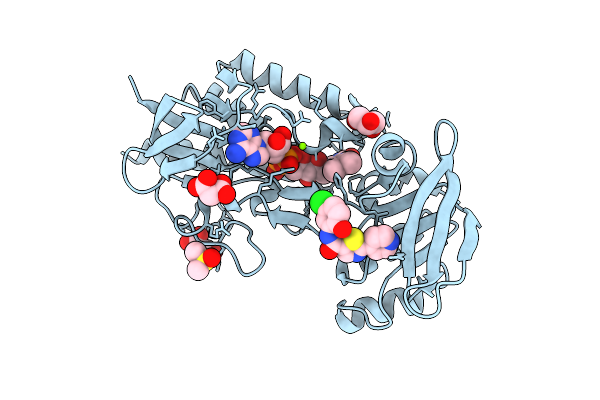 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z1280094148
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I95, DMS |
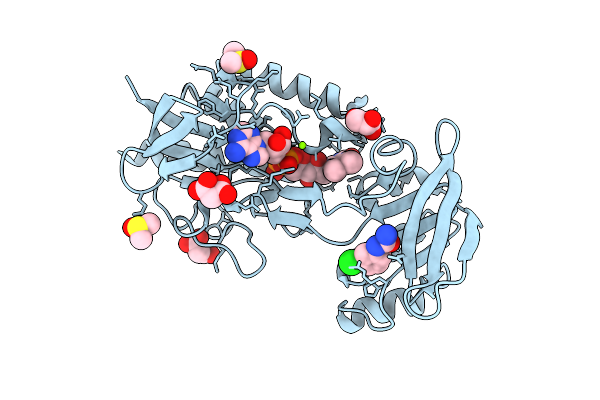 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z166687084
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, DMS, A1I98 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z741560256
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I99 |
 |
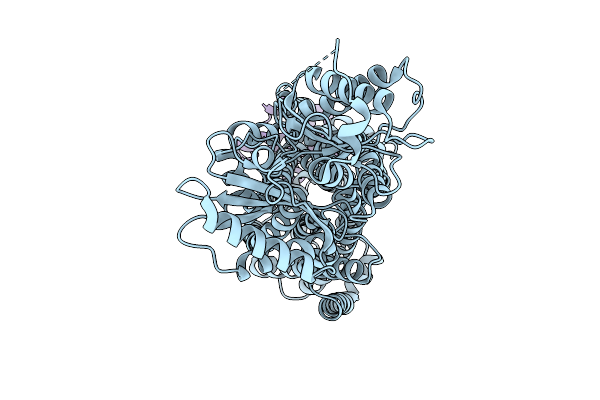
Cryo-Em Map Of Msmeg_3496 In Complex With Acpm, Size Exclusion Chromatography Peak1
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Map Of Msmeg_3496 In Complex With Acpm, Size Exclusion Chromatography Peak2
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN |
 |
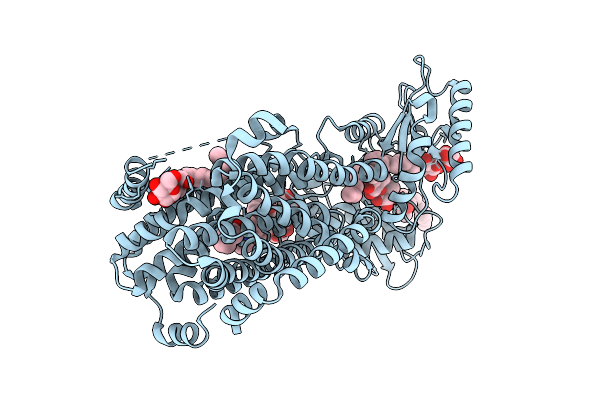
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv), Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN |
 |
Crystal Structure Of Mycolic Acid Transporter Mmpl3 From Mycobacterium Smegmatis Complexed With Indolcarboxamide
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-11-05 Classification: MEMBRANE PROTEIN Ligands: A1L5O, LMT |
 |
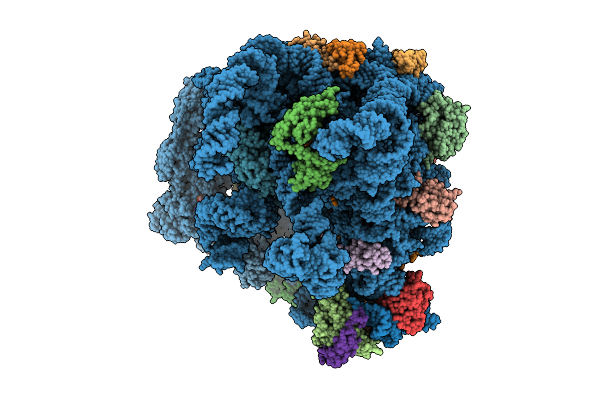
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: RIBOSOME Ligands: PHE, GNP, MG, ZN |
 |
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: RIBOSOME Ligands: GNP, MG, ZN |
 |
Cryo-Em Structure Of The Efflux Transporter Mmpl5/Mmps5 From Mycobacterium Tuberculosis, C3 Symmetry
Organism: Mycobacterium tuberculosis h37rv, Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: MEMBRANE PROTEIN Ligands: CDL, PEV, PN7 |
 |
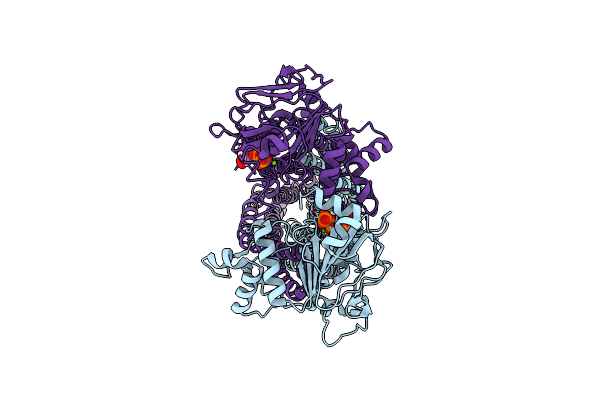
Cryo-Em Structure Of The Efflux Transporter Mmpl5/Mmps5 From Mycobacterium Tuberculosis, C1 Symmetry
Organism: Mycobacterium tuberculosis h37rv, Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-21 Classification: MEMBRANE PROTEIN Ligands: CDL, PEV, PN7 |
 |
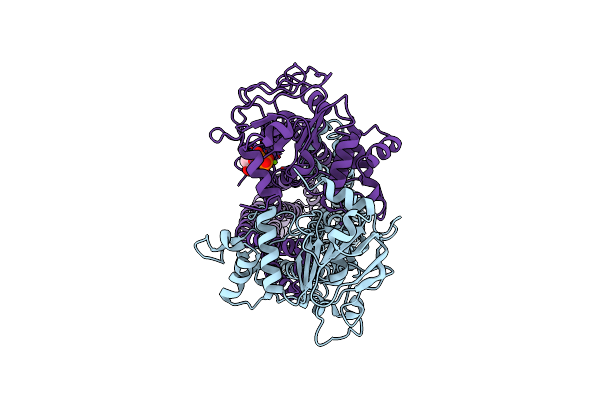
Cryo-Em Structure Of Msrv1273C/72C From Mycobacterium Smegmatis In The Atp|Adp-Bound Ifasym-2 State
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: ATP, MG, ADP |
 |
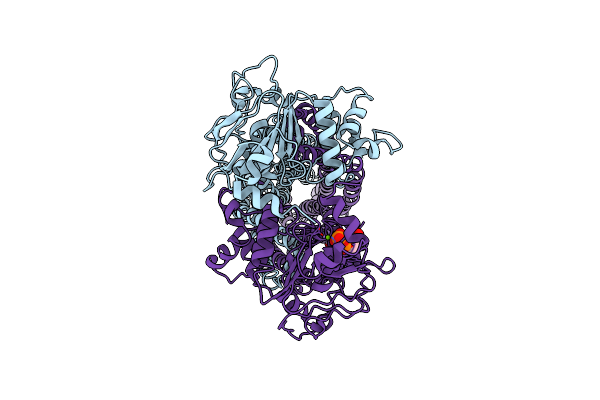
Cryo-Em Structure Of Msrv1273C/72C From Mycobacterium Smegmatis In The Adp-Bound Ifasym-3 State (Atp 37 Degrees C Treated
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: ADP, MG |
 |
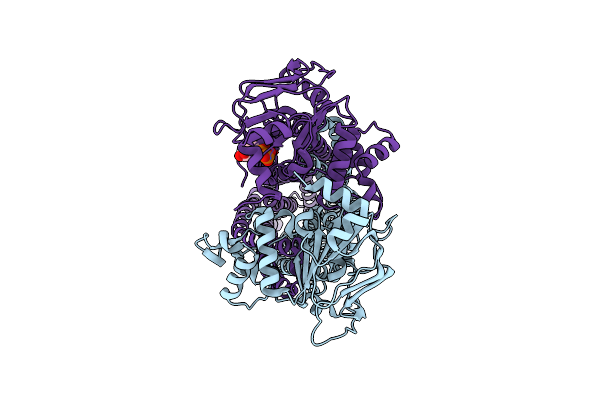
Cryo-Em Structure Of Msrv1273C/72C From Mycobacterium Smegmatis In The Adp-Bound Ifasym-3 State (Adp 4 Degrees C Treated)
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: ADP, MG |
 |
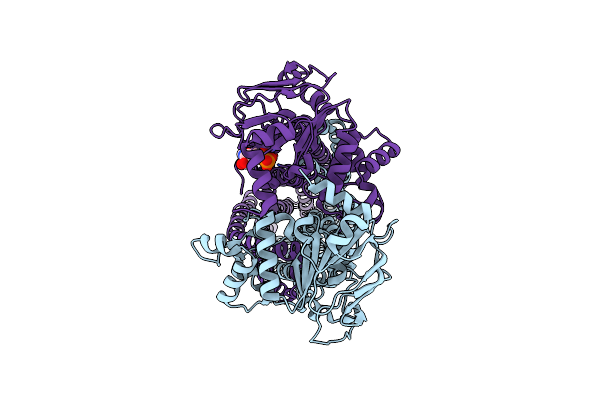
Cryo-Em Structure Of Msrv1273C/72C From Mycobacterium Smegmatis In The Adp-Bound Ifasym-3 (Peptidisc) State (Atp 37Degrees C Treated)
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: ADP |
 |
Cryo-Em Structure Of Msrv1273C/72C From Mycobacterium Smegmatis In The Adp-Bound Ifasym-3 (Peptidisc) State (Adp 4Degrees C Treated)
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: ADP |

