Search Count: 14
All
Selected
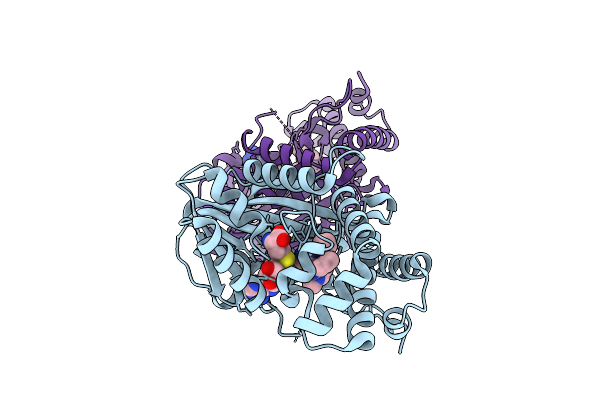 |
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: A1EWC, SAH |
 |
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN |
 |
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: A1EWK, SAH |
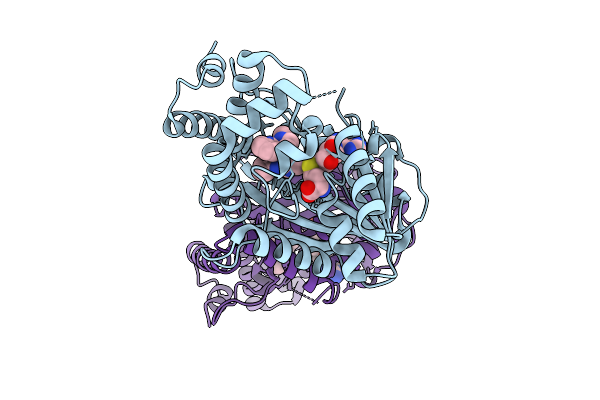 |
N-Methyltransferase 2-Like Complexed With Sah And Ncm In Chimonanthus Praecox
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: SAH, A1EWY |
 |
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: SAH, A1EWY |
 |
N-Methyltransferase 2-Like Complexed With Sah And Mdm In Chimonanthus Praecox
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: A1EW0, SAH |
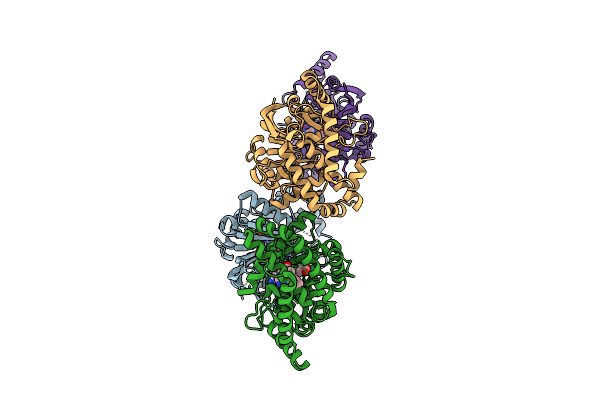 |
N-Methyltransferase 2-Like Complexed With Sah And Pdm In Chimonanthus Praecox
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: SAH, A1EWC |
 |
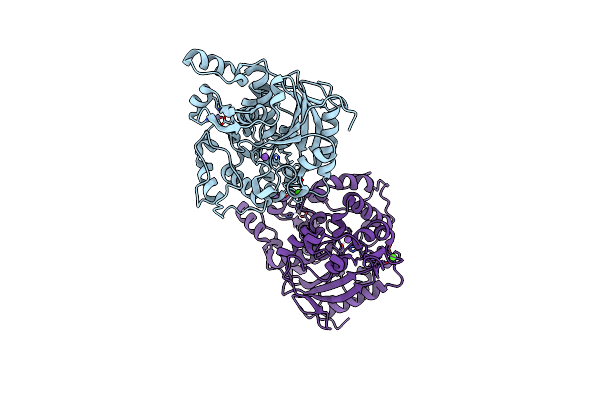
Organism: Chimonanthus praecox
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-01 Classification: PLANT PROTEIN Ligands: SAH, A1EW0 |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2022-12-14 Classification: HYDROLASE Ligands: ZN, CA, NA, CL |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2021-04-07 Classification: HYDROLASE Ligands: ZN, CA |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2019-10-23 Classification: METAL BINDING PROTEIN |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2012-02-22 Classification: HYDROLASE Ligands: ZN, GOL, CL, NA, CA |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2007-10-30 Classification: HYDROLASE Ligands: ZN, CA, CL |
 |
Organism: Geobacillus zalihae
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2007-07-17 Classification: HYDROLASE Ligands: ZN, CA, NA, CL |

